| RAF1 | |||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||
| Identifiers | |||||||||||||||||
| Aliases | RAF1, Raf-1 proto-oncogene, serine/threonine kinase, CMD1NN, CRAF, NS5, Raf-1, c-Raf | ||||||||||||||||
| External IDs | |||||||||||||||||
| |||||||||||||||||
| |||||||||||||||||
| |||||||||||||||||
| Orthologs | |||||||||||||||||
| Species | Human | Mouse | |||||||||||||||
| Entrez | |||||||||||||||||
| Ensembl | |||||||||||||||||
| UniProt | |||||||||||||||||
| RefSeq (mRNA) |
| ||||||||||||||||
| RefSeq (protein) |
|
| |||||||||||||||
| Location (UCSC) | n/a | Chr 6: 115.62 – 115.68 Mb | |||||||||||||||
| PubMed search | [2] | [3] | |||||||||||||||
| Wikidata | |||||||||||||||||
| |||||||||||||||||
RAF proto-oncogene serine/threonine-protein kinase, also known as proto-oncogene c-RAF or simply c-Raf or even Raf-1, is an enzyme[4] that in humans is encoded by the RAF1gene.[5][6] The c-Raf protein is part of the ERK1/2 pathway as a MAP kinase kinase kinase (MAP3K) that functions downstream of the Ras subfamily of membrane associated GTPases.[7] C-Raf is a member of the Raf kinase family of serine/threonine-specific protein kinases, from the TKL (Tyrosine-kinase-like) group of kinases.
- 6Role in cancer
Discovery[edit]
The first Raf gene, v-Raf was found in 1983. It was isolated from the murine retrovirus bearing the number 3611. It was soon demonstrated to be capable to transform rodent fibroblasts to cancerous cell lines, so this gene was given the name Virus-induced Rapidly Accelerated Fibrosarcoma (V-RAF).[5] A year later, another transforming gene was found in the avian retrovirus MH2, named v-Mil - that turned out to be highly similar to v-Raf.[8] Researchers were able to demonstrate that these genes encode enzymes that have serine-threonine kinase activity.[9] Normal cellular homologs of v-Raf and v-Mil were soon found in both the mouse and chicken genome (hence the name c-Raf for the normal cellular Raf gene), and it became clear that these too had a role in regulating growth and cell division.[10][11] Now we know that c-Raf is a principal component of the first described mitogen-activated protein kinase (MAPK) pathway: ERK1/2 signaling.[12] It acts as a MAP3 kinase, initiating the entire kinase cascade. Subsequent experiments showed that the normal, cellular Raf genes can also mutate to become oncogenes, by 'overdriving' MEK1/2 and ERK1/2 activity.[13] In fact, vertebrate genomes contain multiple Raf genes. Several years later after the discovery of c-Raf, two further related kinases were described: A-Raf and B-Raf. The latter became the focus of research in recent years, since a large portion of human tumors carry oncogenic 'driver' mutations in the B-Raf gene.[14] These mutations induce an uncontrolled, high activity of Raf enzymes. Thus diagnostic and therapeutic interest in Raf kinases reached a new peak in the recent years.[15]
Structure[edit]
Apr 03, 1998 Requirement of Ras-GTP-Raf Complexes for Activation of Raf-1 by Protein Kinase C By Richard Marais, Yvonne Light, Clive Mason, Hugh Paterson, Michael F. Olson, Christopher J. Marshall Science 03 Apr 1998: 109-112. Ras activation of the Raf kinase: tyrosine kinase recruitment of the MAP kinase cascade. This pathway is a central effector of cellular differentiation in development; moreover, its inappropriate and continuous activation provides a potent promitogenic force and is a very common occurrence in human cancers. Raf is the best characterized Ras effector and is a member of a family of serine/threonine kinases, that includes Raf-1, A-Raf and B-Raf. Raf activation stimulates a signaling cascade by phosphorylation of MAPK which successively phosphorylate and activate downstream proteins such as ERK1 and ERK2 ( figure 1 figure 1 ). Although activation of Raf-1 is initiated by association with RasGTP on the cytoplasmic membrane, binding of RasGTP to Raf-1 does not induce Raf-1 activation in vitro. Several studies suggested that an additional factor(s) in the plasma membrane is required for Ras-induced activation of Raf-1. Change and the exchange of GDP for GTP. Following Ras activation, Raf is recruited to the cell membrane through Division of Medical Oncology Mayo Clinic, Rochester, MN Submitted for publication Accepted for publication Correspondence to: Julian R. Molina, MD., Ph.D Division of Medical Oncology Mayo Clinic Rochester, MN 55905 Phone: 507- 284 2511.
The human c-Raf gene is located on chromosome 3. At least two isoforms of mRNA have been described (arising from inclusion or removal of an alternative exon) that display only minute differences. The shorter, major isoform - consisting of 17 exons - encodes a protein kinase of 648 amino acids.[16]
A schematic architecture of human c-Raf protein
Similarly to many other MAPKKKs, c-Raf is a multidomain protein, with several additional domains to aid the regulation of its catalytic activity. On its N-terminal segment, a Ras-binding domain (RBD) and a C-kinase homologous domain 1 (C1 domain) are found next to each other. Structures of both conserved domains were solved in the past decades, shedding light on the mechanisms of their regulation.
The Ras-binding domain displays a ubiquitin-like fold (like many other small G-protein associating domains) and selectively binds GTP-bound Ras proteins only.[17][18][19] (You can see this interaction in high detail in the PDB box attached to the article. It shows Rap1 in complex with the RBD of c-Raf.)
The C1 domain - immediately downstream of the Ras binding domain - is a special zinc finger, rich in cysteines and stabilized by two zinc ions. It is similar to the diacylglycerol-binding C1 domains of protein kinase C (PKC) enzymes.[20][21] But unlike PKC, the C1 domains of Raf family kinases do not bind diacylglycerol.[22] Instead, they interact with other lipids, such as ceramide[22] or phosphatidic acid,[23] and even aid in the recognition of activated Ras (GTP-Ras).[21][24]
The close proximity of these two domains as well as several lines of experimental data suggest that they act as a single unit to negatively regulate the activity of the protein kinase domain, by direct physical interaction.[25] Historically, this autoinhibitory block was labelled as the CR1 region ('Conserved Region 1'), the hinge region being named CR2, and the kinase domain CR3. Unfortunately, the precise structure of the autoinhibited kinase remains unknown.
Between the autoinhibitory domain block and the catalytic kinase domain, a long segment - characteristic to all Raf proteins - can be found. It is highly enriched in serine amino acids, but its precise sequence is poorly conserved across related Raf genes. This region appears to be intrinsically unstructured, and very flexible. Its most likely role is to act as a natural 'hinge' between the rigidly folded autoinhibitory and catalytic domains, enabling complex movements and profound conformational rearrangements within the molecule.[26] This hinge region contains a small, conserved island of amino acids, that are responsible for 14-3-3 protein recognition, but only when a critical serine (Ser259 in human c-Raf) is phosphorylated. A second, similar motif is found on the extreme C-terminus (centered around the phosphorylatable Ser 621) of all Raf enzymes, but downstream of the kinase domain.
The C-terminal half of c-Raf folds into a single protein domain, responsible for catalytic activity. The structure of this kinase domain is well-known from both c-Raf[27] and B-Raf.[28] It is highly similar to other Raf kinases and KSR proteins, and distinctly similar to some other MAP3 kinases, such as the Mixed Lineage Kinase (MLK) family. Together they comprise the Tyrosine Kinase Like (TKL) group of protein kinases. Although some features unite their catalytic domains with protein tyrosine kinases, the activity of TKLs is restricted to the phosphorylation of serine and threonine residues within target proteins. The most important substrate of Raf kinases (apart from itself) are the MKK1 and MKK2 kinases, whose activity strictly depends on phosphorylation events performed by Rafs.
Evolutionary relationships[edit]
Human c-Raf is a member of a larger family of related protein kinases. Two further members - found in most vertebrates - belong to the same family: B-Raf and A-Raf. Apart from the different length of their non-conserved N- and C-terminal ends, they all share the same domain architecture, structure and regulation. In comparison to the relatively well-known c-Raf and B-Raf, there is very little known of the precise function of A-Raf, but it is also thought to be similar to the other two members of the family. All these genes are believed to be the product of full gene or genome duplications at the dawn of vertebrate evolution, from a single ancestral Raf gene. Most other animal organisms possess only a single Raf gene. It is called Phl or Draf in Drosophila[29] and Lin-45 in C. elegans.[30]
The family of Raf kinases (schematic architectures)
Multicellular animals also have a type of kinase closely related to Raf: this is the Kinase Suppressor of Ras (KSR). Vertebrates like mammals have two, paralogous KSR genes instead of one: KSR1 and KSR2. Their C-terminal kinase domain is very similar to Raf (originally called CA5 in KSR and CR3 in Raf), but the N-terminal regulatory region differs. Although they also have the flexible hinge (CA4 in KSR) and a C1 domain (CA3 in KSR) before it, KSRs entirely lack the Ras-binding domain. Instead, they have unique regulatory regions on their N-termini, originally termed CA1 ('conserved area 1') and CA2. For a long time, the structure of the CA1 domain was a mystery. However, in 2012, the structure of the CA1 region in KSR1 was solved: it turned out to be a divergent SAM (sterile alpha motif) domain, supplemented with coiled-coils (CC-SAM): this is supposed to aid KSRs in membrane binding.[31] KSRs, like Rafs, also have the twin 14-3-3 associating motifs (that depend on phosphorylation), but also possess novel MAPK-binding motifs on their hinge regions. With a typical sequence Phe-x-Phe-Pro (FxFP) these motifs are important for the feedback regulation of Raf kinases in the ERK1/2 pathway. According to our current knowledge, KSRs also participate in the same pathway as Raf, although they only play an auxiliary role. With a very poor intrinsic kinase activity, they were long thought to be inactive, until their catalytic activity was finally demonstrated in recent years.[32][33] But even then, they contribute only negligibly to MKK1 and MKK2 phosphorylation. The main role of KSR appears to be to provide a heterodimerization partner to Raf enzymes, greatly facilitating their activation by means of allostery. Similar phenomena were described for other MAP3 kinases. ASK2, for example, is a poor enzyme on its own, and it activity appears to be tied to ASK1/ASK2 heterodimerisation.[34]
Raf-like kinases are fully absent from fungi. But recent sequencing of other opisthokonts (e.g. Capsaspora owczarzaki) revealed the presence of genuine Raf kinases in unicellular eukaryotes. Therefore, it is possible that Raf proteins are an ancient heritage and ancestors of fungi secondarily lost Raf-dependent signaling. Fungal MAP kinase pathways that are homologous to the mammalian ERK1/2 pathway (Fus3 and Kss1 in yeast) are activated by MEKK-related kinases (e.g. Ste11 in yeast) instead of Raf enzymes.
Raf kinases found in retroviruses (such as murine v-Raf) are secondarily derived from the corresponding vertebrate genes of their hosts. These Raf genes encode severely truncated proteins, that lack the entire N-terminal autoinhibitory domain, and the 14-3-3 binding motifs. Such severe truncations are known to induce an uncontrolled activity of Raf kinases: that is just exactly what a virus may need for efficient reproduction.
Regulation of activity[edit]
Artist's impression of the autoinhibited state of c-Raf, reinforced by the associated 14-3-3 protein dimers, bound to the phosphorylated twin motifs.[35][36]
As mentioned above, the regulation of c-Raf activity is complex. As a 'gatekeeper' of the ERK1/2 pathway, it is kept in check by a multitude of inhibitory mechanisms, and normally cannot be activated in a single step. The most important regulatory mechanism involves the direct, physical association of the N-terminal autoinhibitory block to the kinase domain of c-Raf. It results in the occlusion of the catalytic site and full shutdown of kinase activity.[25] This 'closed' state can only be relieved if the autoinhibitory domain of Raf engages a partner competing with its own kinase domain, most importantly GTP-bound Ras. Activated small G-proteins can thus break up the intramolacular interactions: this results in a conformational change ('opening') of c-Raf[37] necessary for kinase activation and substrate binding.
14-3-3 proteins also contribute to the autoinhibition. As 14-3-3 proteins are all known to form constitutive dimers, their assemblies have two binding sites.[38] Thus the dimer acts as a 'molecular handcuff', locking their binding partners at a fixed distance and orientation. When the precisely positioned twin 14-3-3 binding motifs are engaged by a single 14-3-3 protein dimer (such as 14-3-3 zeta), they become locked into a conformation that promotes autoinhibition and does not allow the disengagement of the autoinhibitory and catalytic domains.[39] This 'lockdown' of c-Raf (and other Rafs as well as KSRs) is controlled by motif phosphorylation. Unphosphorylated 14-3-3 associating motifs do not bind their partners: they need to get phosphorylated on conserved serines (Ser 259 and Ser 621) first, by other protein kinases. The most important kinase implicated in this event is TGF-beta activated kinase 1 (TAK1), and the enzymes dedicated for removal of these phosphates are the protein phosphatase 1 (PP1) and protein phosphatase 2A (PP2A) complexes.[40][41]
Note that 14-3-3 binding of Raf enzymes is not necessarily inhibitory: once Raf is open and dimerizes, 14-3-3s can also bind in trans, bridging two kinases and 'handcuffing' them together to reinforce the dimer, instead of keeping them away from each other.[42] Further modes of 14-3-3 interactions with c-Raf also exist, but their role is not well known.[43]
Dimerisation is another important mechanism for c-Raf activity regulation and required for Raf activation loop phosphorylation. Normally, only the 'open' kinase domains participate in dimerisation. Unlike B-Raf, that readily forms homodimers with itself, c-Raf prefers heterodimerisation with either B-Raf or KSR1.[citation needed] Homodimers and heterodimers all behave similarly.[33] The B-Raf homodimer kinase domain structure clearly shows that the activation loops (that control the catalytic activity of all known protein kinases) are positioned in an active-like conformation in the dimer. This is due to an allosteric effect of the other molecule binding to the 'back' side of the kinase; such dimers are symmetric and have two, partially active catalytic sites. At this stage, the activity of Raf kinases is low, and unstable.
The activation cycle of mammalian Raf proteins, exemplified by B-Raf (a greatly simplified overview, not showing all steps).[35][36]
To achieve full activity and stabilize the active state, the activation loop of c-Raf needs to be phosphorylated. The only kinases currently known to perform this act are the Raf family kinases themselves. But some other kinases, such as PAK1 can phosphorylate other residues near the kinase domain of c-Raf: the precise role of these auxiliary kinases is unknown. In the context of c-Raf, both c-Raf and KSR1 are needed for the 'transphosphorylation' step. Due to the architecture of the dimers, this phosphorylation can only take place in trans (i.e. one dimer phosphorylates another, in a four-membered transitional complex).[44] By interacting with conserved Arg and Lys residues in the kinase domain, the phosphorylated activation loops shift conformation and become ordered, permanently locking the kinase domain into a fully active state until dephosphorylated. The phosphorylated activation loops also render the kinase insensitive to the presence of its autoinhibitory domain.[45] KSRs cannot undergo this last step as they miss any phosphorylatable residues in their activation loops. But once c-Raf is fully activated, there is no further need to do so: active Raf enzymes can now engage their substrates.[46] Like most protein kinases, c-Raf has multiple substrates. BAD (Bcl2-atagonist of cell death) is directly phosphorylated by c-Raf,[47] along with several types of adenylate cyclases,[48]myosin phosphatase (MYPT),[49]cardiac muscle troponin T (TnTc),[50] etc. The retinoblastoma protein (pRb) and Cdc25 phosphatase were also suggested as possible substrates.[51]
The most important targets of all Raf enzymes are MKK1(MEK1) and MKK2(MEK2). Although the structure of the enzyme-substrate complex c-Raf:MKK1 is unknown, it can be precisely modelled after the KSR2:MKK1 complex.[33] Here no actual catalysis takes place, but it is thought to be highly similar to the way Raf binds to its substrates. The main interaction interface is provided by the C-terminal lobes of both kinase domains; the large, disordered, proline-rich loop unique to MKK1 and MKK2 also plays an important role in its positioning to Raf (and KSR).[52] These MKKs become phosphorylated at at least two sites on their activation loops upon binding to Raf: this will activate them too. The targets of the kinase cascade are ERK1 and ERK2, that are selectively activated by MKK1 or MKK2. ERKs have numerous substrates in cells; they are also capable of translocating into the nucleus to activate nuclear transcription factors. Activated ERKs are pleiotropic effectors of cell physiology and play an important role in the control of gene expression involved in the cell division cycle, cell migration, inhibition of apoptosis, and cell differentiation.
Associated human diseases[edit]
Hereditary gain-of-function mutations of c-Raf are implicated in some rare, but severe syndromes. Most of these mutations involve single amino acid changes at one of the two 14-3-3 binding motifs.[53][54] Mutation of c-Raf is one of the possible causes of Noonan syndrome: affected individuals have congenital heart defects, short and dysmorphic stature and several other deformities. Similar mutations in c-Raf can also cause a related condition, termed LEOPARD syndrome (Lentigo, Electrocardiographic abnormalities, Ocular hypertelorism, Pulmonary stenosis, Abnormal genitalia, Retarded growth, Deafness), with a complex association of defects.
Role in cancer[edit]
Although c-Raf is very clearly capable of mutating into an oncogene in experimental settings, and even in a few human tumors,[55][56] its sister kinase B-Raf is the true major player in carcinogenesis in humans.[57]
B-Raf mutations[edit]
Approximately 20% of all examined human tumor samples display a mutated B-Raf gene.[58] The overwhelming majority of these mutations involve the exchange of a single amino acid: Val 600 into Glu,and this aberrant gene product (BRAF-V600E) can be visualized by immunohistochemistry for clinical molecular diagnostics[59][60] The aberration can mimic the activation loop phosphorylation and - by jumping all control steps at normal activation - immediately render the kinase domain fully active.[61] Since B-Raf can also activate itself by homodimerisation and c-Raf by heterodimerisation, this mutation has a catastrophic effect by turning the ERK1/2 pathway constitutively active, and driving an uncontrolled process of cell division.[62]
As a therapeutic target[edit]
Due to the importance of both Ras and B-Raf mutations in tumorigenesis, several Raf inhibitors were developed to combat cancer, especially against B-Raf exhibiting the V600E mutation. Sorafenib was the first clinically useful agent, that provides a pharmacological alternative to treat previously largely untreatable malignacies, such as renal cell carcinoma and melanoma.[63] Several other molecules followed up, such as Vemurafenib, Regorafenib, Dabrafenib, etc.
Unfortunately, ATP-competitive B-Raf inhibitors may have an undesired effect in K-Ras-dependent cancers: They are simply too selective for B-Raf. While they perfectly well inhibit B-Raf activity in case a mutant B-Raf is the primary culprit, they also promote homo- and heterodimerisation of B-Raf, with itself and c-Raf. This will actually enhance c-Raf activation instead of inhibiting it in case there is no mutation in any Raf genes, but their common upstream activator K-Ras protein is the one mutated.[27] This 'paradoxical' c-Raf activation necessitates the need to screen for B-Raf mutations in patients (by genetic diagnostics) before starting a B-Raf-inhibitor therapy.[64]
List of interacting proteins[edit]
C-Raf has been shown to interact with:
- AKT1,[65]
- ASK1,[66]
- BAG1,[67]
- BRAF,[68]
- Bcl-2,[69]
- CDC25A,[70][71]
- CFLAR,[72]
- FYN,[73]
- GRB10,[74][75]
- HRAS,[76][77][78][79][80][81][82][83][84][85][86][87][88][89][90][91][92]
- HSP90AA1,[93][94]
- KRAS,[81][82]
- MAP2K1,[95]
- MAP3K1,[96]
- MAPK7,[97]
- MAPK8IP3,[98][99]
- PAK1,[100]
- PEBP1,[95]
- PHB,[101]
- PRKCZ,[102]
- RAP1A,[17][86][103][104]
- RHEB,[105][106][107]
- RRAS2[81][108]
- RB1,[101][109]
- RBL2,[109]
- SHOC2,[81]
- STUB1,[93]
- Src,[73]
- TSC22D3,[110]
- YWHAB,[80][102][111][112][113][114]
- YWHAE,[113][114]
- YWHAG,[102][115][116]
- YWHAH,[102][113][117]
- YWHAQ,[95][102][115][118] and
- YWHAZ.[102][119][120][121][122]
See also[edit]
References[edit]
- ^ abcGRCm38: Ensembl release 89: ENSMUSG00000000441 - Ensembl, May 2017
- ^'Human PubMed Reference:'.
- ^'Mouse PubMed Reference:'.
- ^Li P, Wood K, Mamon H, Haser W, Roberts T (February 1991). 'Raf-1: a kinase currently without a cause but not lacking in effects'. Cell. 64 (3): 479–82. doi:10.1016/0092-8674(91)90228-Q. PMID1846778.
- ^ abRapp UR, Goldsborough MD, Mark GE, Bonner TI, Groffen J, Reynolds FH, Stephenson JR (July 1983). 'Structure and biological activity of v-raf, a unique oncogene transduced by a retrovirus'. Proc. Natl. Acad. Sci. U.S.A. 80 (14): 4218–22. Bibcode:1983PNAS..80.4218R. doi:10.1073/pnas.80.14.4218. PMC384008. PMID6308607.
- ^Bonner T, O'Brien SJ, Nash WG, Rapp UR, Morton CC, Leder P (January 1984). 'The human homologs of the raf (mil) oncogene are located on human chromosomes 3 and 4'. Science. 223 (4631): 71–4. Bibcode:1984Sci..223..71B. doi:10.1126/science.6691137. PMID6691137.
- ^'Entrez Gene: RAF1 v-raf-1 murine leukemia viral oncogene homolog 1'.
- ^Sutrave P, Bonner TI, Rapp UR, Jansen HW, Patschinsky T, Bister K (1984). 'Nucleotide sequence of avian retroviral oncogene v-mil: homologue of murine retroviral oncogene v-raf'. Nature. 309 (5963): 85–8. Bibcode:1984Natur.309..85S. doi:10.1038/309085a0. PMID6325930.
- ^Moelling K, Heimann B, Beimling P, Rapp UR, Sander T (1984). 'Serine- and threonine-specific protein kinase activities of purified gag-mil and gag-raf proteins'. Nature. 312 (5994): 558–61. Bibcode:1984Natur.312.558M. doi:10.1038/312558a0. PMID6438534.
- ^Kolch W, Heidecker G, Lloyd P, Rapp UR (January 1991). 'Raf-1 protein kinase is required for growth of induced NIH/3T3 cells'. Nature. 349 (6308): 426–8. Bibcode:1991Natur.349.426K. doi:10.1038/349426a0. PMID1992343.
- ^Mark GE, Rapp UR (April 1984). 'Primary structure of v-raf: relatedness to the src family of oncogenes'. Science. 224 (4646): 285–9. Bibcode:1984Sci..224.285M. doi:10.1126/science.6324342. PMID6324342.
- ^Kyriakis JM, App H, Zhang XF, Banerjee P, Brautigan DL, Rapp UR, Avruch J (July 1992). 'Raf-1 activates MAP kinase-kinase'. Nature. 358 (6385): 417–21. Bibcode:1992Natur.358.417K. doi:10.1038/358417a0. PMID1322500.
- ^Shimizu K, Nakatsu Y, Nomoto S, Sekiguchi M (1986). 'Structure of the activated c-raf-1 gene from human stomach cancer'. Int. Symp. Princess Takamatsu Cancer Res. Fund. 17: 85–91. PMID2843497.
- ^Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, Teague J, Woffendin H, Garnett MJ, Bottomley W, Davis N, Dicks E, Ewing R, Floyd Y, Gray K, Hall S, Hawes R, Hughes J, Kosmidou V, Menzies A, Mould C, Parker A, Stevens C, Watt S, Hooper S, Wilson R, Jayatilake H, Gusterson BA, Cooper C, Shipley J, Hargrave D, Pritchard-Jones K, Maitland N, Chenevix-Trench G, Riggins GJ, Bigner DD, Palmieri G, Cossu A, Flanagan A, Nicholson A, Ho JW, Leung SY, Yuen ST, Weber BL, Seigler HF, Darrow TL, Paterson H, Marais R, Marshall CJ, Wooster R, Stratton MR, Futreal PA (June 2002). 'Mutations of the BRAF gene in human cancer'. Nature. 417 (6892): 949–54. doi:10.1038/nature00766. PMID12068308.
- ^Sridhar SS, Hedley D, Siu LL (April 2005). 'Raf kinase as a target for anticancer therapeutics'. Mol. Cancer Ther. 4 (4): 677–85. doi:10.1158/1535-7163.MCT-04-0297. PMID15827342.
- ^Dozier C, Ansieau S, Ferreira E, Coll J, Stehelin D (August 1991). 'An alternatively spliced c-mil/raf mRNA is predominantly expressed in chicken muscular tissues and conserved among vertebrate species'. Oncogene. 6 (8): 1307–11. PMID1886707.
- ^ abNassar N, Horn G, Herrmann C, Scherer A, McCormick F, Wittinghofer A (June 1995). 'The 2.2 A crystal structure of the Ras-binding domain of the serine/threonine kinase c-Raf1 in complex with Rap1A and a GTP analogue'. Nature. 375 (6532): 554–60. Bibcode:1995Natur.375.554N. doi:10.1038/375554a0. PMID7791872.
- ^Emerson SD, Madison VS, Palermo RE, Waugh DS, Scheffler JE, Tsao KL, Kiefer SE, Liu SP, Fry DC (May 1995). 'Solution structure of the Ras-binding domain of c-Raf-1 and identification of its Ras interaction surface'. Biochemistry. 34 (21): 6911–8. doi:10.1021/bi00021a001. PMID7766599.
- ^Moodie SA, Willumsen BM, Weber MJ, Wolfman A (June 1993). 'Complexes of Ras.GTP with Raf-1 and mitogen-activated protein kinase kinase'. Science. 260 (5114): 1658–61. Bibcode:1993Sci..260.1658M. doi:10.1126/science.8503013. PMID8503013.
- ^Mott HR, Carpenter JW, Zhong S, Ghosh S, Bell RM, Campbell SL (August 1996). 'The solution structure of the Raf-1 cysteine-rich domain: a novel ras and phospholipid binding site'. Proc. Natl. Acad. Sci. U.S.A. 93 (16): 8312–7. Bibcode:1996PNAS..93.8312M. doi:10.1073/pnas.93.16.8312. PMC38667. PMID8710867.
- ^ abDaub M, Jöckel J, Quack T, Weber CK, Schmitz F, Rapp UR, Wittinghofer A, Block C (November 1998). 'The RafC1 cysteine-rich domain contains multiple distinct regulatory epitopes which control Ras-dependent Raf activation'. Mol. Cell. Biol. 18 (11): 6698–710. PMC109253. PMID9774683.
- ^ abYin X, Zafrullah M, Lee H, Haimovitz-Friedman A, Fuks Z, Kolesnick R (2009). 'A ceramide-binding C1 domain mediates kinase suppressor of ras membrane translocation'. Cell. Physiol. Biochem. 24 (3–4): 219–30. doi:10.1159/000233248. PMC2978518. PMID19710537.
- ^Kraft CA, Garrido JL, Fluharty E, Leiva-Vega L, Romero G (December 2008). 'Role of phosphatidic acid in the coupling of the ERK cascade'. J. Biol. Chem. 283 (52): 36636–45. doi:10.1074/jbc.M804633200. PMC2606017. PMID18952605.
- ^Brtva TR, Drugan JK, Ghosh S, Terrell RS, Campbell-Burk S, Bell RM, Der CJ (April 1995). 'Two distinct Raf domains mediate interaction with Ras'. J. Biol. Chem. 270 (17): 9809–12. doi:10.1074/jbc.270.17.9809. PMID7730360.
- ^ abCutler RE, Stephens RM, Saracino MR, Morrison DK (August 1998). 'Autoregulation of the Raf-1 serine/threonine kinase'. Proc. Natl. Acad. Sci. U.S.A. 95 (16): 9214–9. Bibcode:1998PNAS..95.9214C. doi:10.1073/pnas.95.16.9214. PMC21318. PMID9689060.
- ^Hmitou I, Druillennec S, Valluet A, Peyssonnaux C, Eychène A (January 2007). 'Differential regulation of B-raf isoforms by phosphorylation and autoinhibitory mechanisms'. Mol. Cell. Biol. 27 (1): 31–43. doi:10.1128/MCB.01265-06. PMC1800654. PMID17074813.
- ^ abHatzivassiliou G, Song K, Yen I, Brandhuber BJ, Anderson DJ, Alvarado R, Ludlam MJ, Stokoe D, Gloor SL, Vigers G, Morales T, Aliagas I, Liu B, Sideris S, Hoeflich KP, Jaiswal BS, Seshagiri S, Koeppen H, Belvin M, Friedman LS, Malek S (March 2010). 'RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth'. Nature. 464 (7287): 431–5. Bibcode:2010Natur.464.431H. doi:10.1038/nature08833. PMID20130576.
- ^Wan PT, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D, Good VM, Jones CM, Marshall CJ, Springer CJ, Barford D, Marais R (March 2004). 'Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF'. Cell. 116 (6): 855–67. doi:10.1016/S0092-8674(04)00215-6. PMID15035987.
- ^Mark GE, MacIntyre RJ, Digan ME, Ambrosio L, Perrimon N (June 1987). 'Drosophila melanogaster homologs of the raf oncogene'. Mol. Cell. Biol. 7 (6): 2134–40. PMC365335. PMID3037346.
- ^Chong H, Vikis HG, Guan KL (May 2003). 'Mechanisms of regulating the Raf kinase family'. Cell. Signal. 15 (5): 463–9. doi:10.1016/S0898-6568(02)00139-0. PMID12639709.
- ^Koveal D, Schuh-Nuhfer N, Ritt D, Page R, Morrison DK, Peti W (December 2012). 'A CC-SAM, for coiled coil-sterile α motif, domain targets the scaffold KSR-1 to specific sites in the plasma membrane'. Sci Signal. 5 (255): ra94. doi:10.1126/scisignal.2003289. PMC3740349. PMID23250398.
- ^Hu J, Yu H, Kornev AP, Zhao J, Filbert EL, Taylor SS, Shaw AS (April 2011). 'Mutation that blocks ATP binding creates a pseudokinase stabilizing the scaffolding function of kinase suppressor of Ras, CRAF and BRAF'. Proc. Natl. Acad. Sci. U.S.A. 108 (15): 6067–72. Bibcode:2011PNAS.108.6067H. doi:10.1073/pnas.1102554108. PMC3076888. PMID21441104.
- ^ abcBrennan DF, Dar AC, Hertz NT, Chao WC, Burlingame AL, Shokat KM, Barford D (April 2011). 'A Raf-induced allosteric transition of KSR stimulates phosphorylation of MEK'. Nature. 472 (7343): 366–9. Bibcode:2011Natur.472.366B. doi:10.1038/nature09860. PMID21441910.
- ^Ortner E, Moelling K (October 2007). 'Heteromeric complex formation of ASK2 and ASK1 regulates stress-induced signaling'. Biochem. Biophys. Res. Commun. 362 (2): 454–9. doi:10.1016/j.bbrc.2007.08.006. PMID17714688.
- ^ abMatallanas D, Birtwistle M, Romano D, Zebisch A, Rauch J, von Kriegsheim A, Kolch W (2011). 'Raf family kinases: old dogs have learned new tricks'. Genes Cancer. 2 (3): 232–60. doi:10.1177/1947601911407323. PMC3128629. PMID21779496.
- ^ abAlexa A, Varga J, Reményi A (2010). 'Scaffolds are 'active' regulators of signaling modules'. FEBS J. 277 (21): 4376–82. doi:10.1111/j.1742-4658.2010.07867.x. PMID20883493.
- ^Terai K, Matsuda M (March 2005). 'Ras binding opens c-Raf to expose the docking site for mitogen-activated protein kinase kinase'. EMBO Rep. 6 (3): 251–5. doi:10.1038/sj.embor.7400349. PMC1299259. PMID15711535.
- ^Liu D, Bienkowska J, Petosa C, Collier RJ, Fu H, Liddington R (July 1995). 'Crystal structure of the zeta isoform of the 14-3-3 protein'. Nature. 376 (6536): 191–4. Bibcode:1995Natur.376.191L. doi:10.1038/376191a0. PMID7603574.
- ^Fischer A, Baljuls A, Reinders J, Nekhoroshkova E, Sibilski C, Metz R, Albert S, Rajalingam K, Hekman M, Rapp UR (January 2009). 'Regulation of RAF activity by 14-3-3 proteins: RAF kinases associate functionally with both homo- and heterodimeric forms of 14-3-3 proteins'. J. Biol. Chem. 284 (5): 3183–94. doi:10.1074/jbc.M804795200. PMID19049963.
- ^Rodriguez-Viciana P, Oses-Prieto J, Burlingame A, Fried M, McCormick F (April 2006). 'A phosphatase holoenzyme comprised of Shoc2/Sur8 and the catalytic subunit of PP1 functions as an M-Ras effector to modulate Raf activity'. Mol. Cell. 22 (2): 217–30. doi:10.1016/j.molcel.2006.03.027. PMID16630891.
- ^Jaumot M, Hancock JF (July 2001). 'Protein phosphatases 1 and 2A promote Raf-1 activation by regulating 14-3-3 interactions'. Oncogene. 20 (30): 3949–58. doi:10.1038/sj.onc.1204526. PMID11494123.
- ^Tzivion G, Luo Z, Avruch J (July 1998). 'A dimeric 14-3-3 protein is an essential cofactor for Raf kinase activity'. Nature. 394 (6688): 88–92. Bibcode:1998Natur.394..88T. doi:10.1038/27938. PMID9665134.
- ^Molzan M, Ottmann C (November 2012). 'Synergistic binding of the phosphorylated S233- and S259-binding sites of C-RAF to one 14-3-3ζ dimer'. J. Mol. Biol. 423 (4): 486–95. doi:10.1016/j.jmb.2012.08.009. PMID22922483.
- ^McKay MM, Freeman AK, Morrison DK (2011). 'Complexity in KSR function revealed by Raf inhibitor and KSR structure studies'. Small GTPases. 2 (5): 276–281. doi:10.4161/sgtp.2.5.17740. PMC3265819. PMID22292131.
- ^Chong H, Guan KL (September 2003). 'Regulation of Raf through phosphorylation and N terminus-C terminus interaction'. J. Biol. Chem. 278 (38): 36269–76. doi:10.1074/jbc.M212803200. PMID12865432.
- ^Shi F, Lemmon MA (May 2011). 'Biochemistry. KSR plays CRAF-ty'. Science. 332 (6033): 1043–4. Bibcode:2011Sci..332.1043S. doi:10.1126/science.1208063. PMID21617065.
- ^Ye DZ, Jin S, Zhuo Y, Field J (2011). Bauer JA (ed.). 'p21-Activated kinase 1 (Pak1) phosphorylates BAD directly at serine 111 in vitro and indirectly through Raf-1 at serine 112'. PLoS ONE. 6 (11): e27637. Bibcode:2011PLoSO..627637Y. doi:10.1371/journal.pone.0027637. PMC3214075. PMID22096607.
- ^Ding Q, Gros R, Gray ID, Taussig R, Ferguson SS, Feldman RD (October 2004). 'Raf kinase activation of adenylyl cyclases: isoform-selective regulation'. Mol. Pharmacol. 66 (4): 921–8. doi:10.1124/mol.66.4.921. PMID15385642.
- ^Broustas CG, Grammatikakis N, Eto M, Dent P, Brautigan DL, Kasid U (January 2002). 'Phosphorylation of the myosin-binding subunit of myosin phosphatase by Raf-1 and inhibition of phosphatase activity'. J. Biol. Chem. 277 (4): 3053–9. doi:10.1074/jbc.M106343200. PMID11719507.
- ^Pfleiderer P, Sumandea MP, Rybin VO, Wang C, Steinberg SF (2009). 'Raf-1: a novel cardiac troponin T kinase'. J. Muscle Res. Cell. Motil. 30 (1–2): 67–72. doi:10.1007/s10974-009-9176-y. PMC2893395. PMID19381846.
- ^Hindley A, Kolch W (April 2002). 'Extracellular signal regulated kinase (ERK)/mitogen activated protein kinase (MAPK)-independent functions of Raf kinases'. J. Cell Sci. 115 (Pt 8): 1575–81. PMID11950876.
- ^Catling AD, Schaeffer HJ, Reuter CW, Reddy GR, Weber MJ (October 1995). 'A proline-rich sequence unique to MEK1 and MEK2 is required for raf binding and regulates MEK function'. Mol. Cell. Biol. 15 (10): 5214–25. PMC230769. PMID7565670.
- ^Pandit B, Sarkozy A, Pennacchio LA, Carta C, Oishi K, Martinelli S, Pogna EA, Schackwitz W, Ustaszewska A, Landstrom A, Bos JM, Ommen SR, Esposito G, Lepri F, Faul C, Mundel P, López Siguero JP, Tenconi R, Selicorni A, Rossi C, Mazzanti L, Torrente I, Marino B, Digilio MC, Zampino G, Ackerman MJ, Dallapiccola B, Tartaglia M, Gelb BD (August 2007). 'Gain-of-function RAF1 mutations cause Noonan and LEOPARD syndromes with hypertrophic cardiomyopathy'. Nat. Genet. 39 (8): 1007–12. doi:10.1038/ng2073. PMID17603483.
- ^Molzan M, Schumacher B, Ottmann C, Baljuls A, Polzien L, Weyand M, Thiel P, Rose R, Rose M, Kuhenne P, Kaiser M, Rapp UR, Kuhlmann J, Ottmann C (October 2010). 'Impaired binding of 14-3-3 to C-RAF in Noonan syndrome suggests new approaches in diseases with increased Ras signaling'. Mol. Cell. Biol. 30 (19): 4698–711. doi:10.1128/MCB.01636-09. PMC2950525. PMID20679480.
- ^Storm SM, Rapp UR (April 1993). 'Oncogene activation: c-raf-1 gene mutations in experimental and naturally occurring tumors'. Toxicol. Lett. 67 (1–3): 201–10. doi:10.1016/0378-4274(93)90056-4. PMID8451761.
- ^Zebisch A, Staber PB, Delavar A, Bodner C, Hiden K, Fischereder K, Janakiraman M, Linkesch W, Auner HW, Emberger W, Windpassinger C, Schimek MG, Hoefler G, Troppmair J, Sill H (April 2006). 'Two transforming C-RAF germ-line mutations identified in patients with therapy-related acute myeloid leukemia'. Cancer Res. 66 (7): 3401–8. doi:10.1158/0008-5472.CAN-05-0115. PMID16585161.
- ^Emuss V, Garnett M, Mason C, Marais R (November 2005). 'Mutations of C-RAF are rare in human cancer because C-RAF has a low basal kinase activity compared with B-RAF'. Cancer Res. 65 (21): 9719–26. doi:10.1158/0008-5472.CAN-05-1683. PMID16266992.
- ^Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D, Jia M, Shepherd R, Leung K, Menzies A, Teague JW, Campbell PJ, Stratton MR, Futreal PA (January 2011). 'COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer'. Nucleic Acids Res. 39 (Database issue): D945–50. doi:10.1093/nar/gkq929. PMC3013785. PMID20952405.
- ^Capper D, Berghoff AS, Magerle M, Ilhan A, Wöhrer A, Hackl M, Pichler J, Pusch S, Meyer J, Habel A, Petzelbauer P, Birner P, von Deimling A, Preusser M (2012). 'Immunohistochemical testing of BRAF V600E status in 1,120 tumor tissue samples of patients with brain metastases'. Acta Neuropathol. 123 (2): 223–33. doi:10.1007/s00401-011-0887-y. PMID22012135.
- ^Capper D, Preusser M, Habel A, Sahm F, Ackermann U, Schindler G, Pusch S, Mechtersheimer G, Zentgraf H, von Deimling A (2011). 'Assessment of BRAF V600E mutation status by immunohistochemistry with a mutation-specific monoclonal antibody'. Acta Neuropathol. 122 (1): 11–9. doi:10.1007/s00401-011-0841-z. PMID21638088.
- ^Tran NH, Wu X, Frost JA (April 2005). 'B-Raf and Raf-1 are regulated by distinct autoregulatory mechanisms'. J. Biol. Chem. 280 (16): 16244–53. doi:10.1074/jbc.M501185200. PMID15710605.
- ^Garnett MJ, Rana S, Paterson H, Barford D, Marais R (December 2005). 'Wild-type and mutant B-RAF activate C-RAF through distinct mechanisms involving heterodimerization'. Mol. Cell. 20 (6): 963–9. doi:10.1016/j.molcel.2005.10.022. PMID16364920.
- ^Maurer G, Tarkowski B, Baccarini M (August 2011). 'Raf kinases in cancer-roles and therapeutic opportunities'. Oncogene. 30 (32): 3477–88. doi:10.1038/onc.2011.160. PMID21577205.
- ^Kim DH, Sim T (March 2012). 'Novel small molecule Raf kinase inhibitors for targeted cancer therapeutics'. Arch. Pharm. Res. 35 (4): 605–15. doi:10.1007/s12272-012-0403-5. PMID22553052.
- ^Zimmermann S, Moelling K (November 1999). 'Phosphorylation and regulation of Raf by Akt (protein kinase B)'. Science. 286 (5445): 1741–4. doi:10.1126/science.286.5445.1741. PMID10576742.
- ^Chen J, Fujii K, Zhang L, Roberts T, Fu H (July 2001). 'Raf-1 promotes cell survival by antagonizing apoptosis signal-regulating kinase 1 through a MEK-ERK independent mechanism'. Proc. Natl. Acad. Sci. U.S.A. 98 (14): 7783–8. Bibcode:2001PNAS..98.7783C. doi:10.1073/pnas.141224398. PMC35419. PMID11427728.
- ^Wang HG, Takayama S, Rapp UR, Reed JC (July 1996). 'Bcl-2 interacting protein, BAG-1, binds to and activates the kinase Raf-1'. Proc. Natl. Acad. Sci. U.S.A. 93 (14): 7063–8. Bibcode:1996PNAS..93.7063W. doi:10.1073/pnas.93.14.7063. PMC38936. PMID8692945.
- ^Weber CK, Slupsky JR, Kalmes HA, Rapp UR (May 2001). 'Active Ras induces heterodimerization of cRaf and BRaf'. Cancer Res. 61 (9): 3595–8. PMID11325826.
- ^Wang HG, Rapp UR, Reed JC (November 1996). 'Bcl-2 targets the protein kinase Raf-1 to mitochondria'. Cell. 87 (4): 629–38. doi:10.1016/s0092-8674(00)81383-5. PMID8929532.
- ^Galaktionov K, Jessus C, Beach D (May 1995). 'Raf1 interaction with Cdc25 phosphatase ties mitogenic signal transduction to cell cycle activation'. Genes Dev. 9 (9): 1046–58. doi:10.1101/gad.9.9.1046. PMID7744247.
- ^Huang TS, Shu CH, Yang WK, Whang-Peng J (July 1997). 'Activation of CDC 25 phosphatase and CDC 2 kinase involved in GL331-induced apoptosis'. Cancer Res. 57 (14): 2974–8. PMID9230211.
- ^Kataoka T, Budd RC, Holler N, Thome M, Martinon F, Irmler M, Burns K, Hahne M, Kennedy N, Kovacsovics M, Tschopp J (June 2000). 'The caspase-8 inhibitor FLIP promotes activation of NF-kappaB and Erk signaling pathways'. Curr. Biol. 10 (11): 640–8. doi:10.1016/s0960-9822(00)00512-1. PMID10837247.
- ^ abCleghon V, Morrison DK (July 1994). 'Raf-1 interacts with Fyn and Src in a non-phosphotyrosine-dependent manner'. J. Biol. Chem. 269 (26): 17749–55. PMID7517401.
- ^Nantel A, Huber M, Thomas DY (December 1999). 'Localization of endogenous Grb10 to the mitochondria and its interaction with the mitochondrial-associated Raf-1 pool'. J. Biol. Chem. 274 (50): 35719–24. doi:10.1074/jbc.274.50.35719. PMID10585452.
- ^Nantel A, Mohammad-Ali K, Sherk J, Posner BI, Thomas DY (April 1998). 'Interaction of the Grb10 adapter protein with the Raf1 and MEK1 kinases'. J. Biol. Chem. 273 (17): 10475–84. doi:10.1074/jbc.273.17.10475. PMID9553107.
- ^Stang S, Bottorff D, Stone JC (June 1997). 'Interaction of activated Ras with Raf-1 alone may be sufficient for transformation of rat2 cells'. Mol. Cell. Biol. 17 (6): 3047–55. PMC232157. PMID9154803.
- ^Germani A, Prabel A, Mourah S, Podgorniak MP, Di Carlo A, Ehrlich R, Gisselbrecht S, Varin-Blank N, Calvo F, Bruzzoni-Giovanelli H (December 2003). 'SIAH-1 interacts with CtIP and promotes its degradation by the proteasome pathway'. Oncogene. 22 (55): 8845–51. doi:10.1038/sj.onc.1206994. PMID14654780.
- ^Mitin NY, Ramocki MB, Zullo AJ, Der CJ, Konieczny SF, Taparowsky EJ (May 2004). 'Identification and characterization of rain, a novel Ras-interacting protein with a unique subcellular localization'. J. Biol. Chem. 279 (21): 22353–61. doi:10.1074/jbc.M312867200. PMID15031288.
- ^Vargiu P, De Abajo R, Garcia-Ranea JA, Valencia A, Santisteban P, Crespo P, Bernal J (January 2004). 'The small GTP-binding protein, Rhes, regulates signal transduction from G protein-coupled receptors'. Oncogene. 23 (2): 559–68. doi:10.1038/sj.onc.1207161. PMID14724584.
- ^ abYuryev A, Wennogle LP (February 2003). 'Novel raf kinase protein-protein interactions found by an exhaustive yeast two-hybrid analysis'. Genomics. 81 (2): 112–25. doi:10.1016/s0888-7543(02)00008-3. PMID12620389.
- ^ abcdLi W, Han M, Guan KL (April 2000). 'The leucine-rich repeat protein SUR-8 enhances MAP kinase activation and forms a complex with Ras and Raf'. Genes Dev. 14 (8): 895–900. PMC316541. PMID10783161.
- ^ abKiyono M, Kato J, Kataoka T, Kaziro Y, Satoh T (September 2000). 'Stimulation of Ras guanine nucleotide exchange activity of Ras-GRF1/CDC25(Mm) upon tyrosine phosphorylation by the Cdc42-regulated kinase ACK1'. J. Biol. Chem. 275 (38): 29788–93. doi:10.1074/jbc.M001378200. PMID10882715.
- ^Janoueix-Lerosey I, Pasheva E, de Tand MF, Tavitian A, de Gunzburg J (March 1998). 'Identification of a specific effector of the small GTP-binding protein Rap2'. Eur. J. Biochem. 252 (2): 290–8. doi:10.1046/j.1432-1327.1998.2520290.x. PMID9523700.
- ^Boettner B, Govek EE, Cross J, Van Aelst L (August 2000). 'The junctional multidomain protein AF-6 is a binding partner of the Rap1A GTPase and associates with the actin cytoskeletal regulator profilin'. Proc. Natl. Acad. Sci. U.S.A. 97 (16): 9064–9. Bibcode:2000PNAS..97.9064B. doi:10.1073/pnas.97.16.9064. PMC16822. PMID10922060.
- ^Karbowniczek M, Robertson GP, Henske EP (September 2006). 'Rheb inhibits C-raf activity and B-raf/C-raf heterodimerization'. J. Biol. Chem. 281 (35): 25447–56. doi:10.1074/jbc.M605273200. PMID16803888.
- ^ abHan L, Colicelli J (March 1995). 'A human protein selected for interference with Ras function interacts directly with Ras and competes with Raf1'. Mol. Cell. Biol. 15 (3): 1318–23. doi:10.1128/mcb.15.3.1318. PMC230355. PMID7862125.
- ^Jelinek T, Catling AD, Reuter CW, Moodie SA, Wolfman A, Weber MJ (December 1994). 'RAS and RAF-1 form a signalling complex with MEK-1 but not MEK-2'. Mol. Cell. Biol. 14 (12): 8212–8. PMC359360. PMID7969158.
- ^Romero F, Martínez-A C, Camonis J, Rebollo A (June 1999). 'Aiolos transcription factor controls cell death in T cells by regulating Bcl-2 expression and its cellular localization'. EMBO J. 18 (12): 3419–30. doi:10.1093/emboj/18.12.3419. PMC1171421. PMID10369681.
- ^Morcos P, Thapar N, Tusneem N, Stacey D, Tamanoi F (May 1996). 'Identification of neurofibromin mutants that exhibit allele specificity or increased Ras affinity resulting in suppression of activated ras alleles'. Mol. Cell. Biol. 16 (5): 2496–503. PMC231238. PMID8628317.
- ^Hu CD, Kariya K, Tamada M, Akasaka K, Shirouzu M, Yokoyama S, Kataoka T (December 1995). 'Cysteine-rich region of Raf-1 interacts with activator domain of post-translationally modified Ha-Ras'. J. Biol. Chem. 270 (51): 30274–7. doi:10.1074/jbc.270.51.30274. PMID8530446.
- ^Rodriguez-Viciana P, Warne PH, Khwaja A, Marte BM, Pappin D, Das P, Waterfield MD, Ridley A, Downward J (May 1997). 'Role of phosphoinositide 3-OH kinase in cell transformation and control of the actin cytoskeleton by Ras'. Cell. 89 (3): 457–67. doi:10.1016/s0092-8674(00)80226-3. PMID9150145.
- ^Huang YZ, Zang M, Xiong WC, Luo Z, Mei L (January 2003). 'Erbin suppresses the MAP kinase pathway'. J. Biol. Chem. 278 (2): 1108–14. doi:10.1074/jbc.M205413200. PMID12379659.
- ^ abDogan T, Harms GS, Hekman M, Karreman C, Oberoi TK, Alnemri ES, Rapp UR, Rajalingam K (December 2008). 'X-linked and cellular IAPs modulate the stability of C-RAF kinase and cell motility'. Nat. Cell Biol. 10 (12): 1447–55. doi:10.1038/ncb1804. PMID19011619.
- ^Stancato LF, Chow YH, Hutchison KA, Perdew GH, Jove R, Pratt WB (October 1993). 'Raf exists in a native heterocomplex with hsp90 and p50 that can be reconstituted in a cell-free system'. J. Biol. Chem. 268 (29): 21711–6. PMID8408024.
- ^ abcYeung K, Janosch P, McFerran B, Rose DW, Mischak H, Sedivy JM, Kolch W (May 2000). 'Mechanism of suppression of the Raf/MEK/extracellular signal-regulated kinase pathway by the raf kinase inhibitor protein'. Mol. Cell. Biol. 20 (9): 3079–85. doi:10.1128/mcb.20.9.3079-3085.2000. PMC85596. PMID10757792.
- ^Karandikar M, Xu S, Cobb MH (December 2000). 'MEKK1 binds raf-1 and the ERK2 cascade components'. J. Biol. Chem. 275 (51): 40120–7. doi:10.1074/jbc.M005926200. PMID10969079.
- ^English JM, Pearson G, Hockenberry T, Shivakumar L, White MA, Cobb MH (October 1999). 'Contribution of the ERK5/MEK5 pathway to Ras/Raf signaling and growth control'. J. Biol. Chem. 274 (44): 31588–92. doi:10.1074/jbc.274.44.31588. PMID10531364.
- ^Kuboki Y, Ito M, Takamatsu N, Yamamoto KI, Shiba T, Yoshioka K (December 2000). 'A scaffold protein in the c-Jun NH2-terminal kinase signaling pathways suppresses the extracellular signal-regulated kinase signaling pathways'. J. Biol. Chem. 275 (51): 39815–8. doi:10.1074/jbc.C000403200. PMID11044439.
- ^Ito M, Yoshioka K, Akechi M, Yamashita S, Takamatsu N, Sugiyama K, Hibi M, Nakabeppu Y, Shiba T, Yamamoto KI (November 1999). 'JSAP1, a novel jun N-terminal protein kinase (JNK)-binding protein that functions as a Scaffold factor in the JNK signaling pathway'. Mol. Cell. Biol. 19 (11): 7539–48. doi:10.1128/mcb.19.11.7539. PMC84763. PMID10523642.
- ^Zang M, Hayne C, Luo Z (February 2002). 'Interaction between active Pak1 and Raf-1 is necessary for phosphorylation and activation of Raf-1'. J. Biol. Chem. 277 (6): 4395–405. doi:10.1074/jbc.M110000200. PMID11733498.
- ^ abWang S, Nath N, Fusaro G, Chellappan S (November 1999). 'Rb and prohibitin target distinct regions of E2F1 for repression and respond to different upstream signals'. Mol. Cell. Biol. 19 (11): 7447–60. doi:10.1128/mcb.19.11.7447. PMC84738. PMID10523633.
- ^ abcdefVan Der Hoeven PC, Van Der Wal JC, Ruurs P, Van Dijk MC, Van Blitterswijk J (January 2000). '14-3-3 isotypes facilitate coupling of protein kinase C-zeta to Raf-1: negative regulation by 14-3-3 phosphorylation'. Biochem. J. 345 (2): 297–306. doi:10.1042/0264-6021:3450297. PMC1220759. PMID10620507.
- ^Hu CD, Kariya K, Okada T, Qi X, Song C, Kataoka T (January 1999). 'Effect of phosphorylation on activities of Rap1A to interact with Raf-1 and to suppress Ras-dependent Raf-1 activation'. J. Biol. Chem. 274 (1): 48–51. doi:10.1074/jbc.274.1.48. PMID9867809.
- ^Okada T, Hu CD, Jin TG, Kariya K, Yamawaki-Kataoka Y, Kataoka T (September 1999). 'The strength of interaction at the Raf cysteine-rich domain is a critical determinant of response of Raf to Ras family small GTPases'. Mol. Cell. Biol. 19 (9): 6057–64. doi:10.1128/mcb.19.9.6057. PMC84512. PMID10454553.
- ^Long X, Lin Y, Ortiz-Vega S, Yonezawa K, Avruch J (April 2005). 'Rheb binds and regulates the mTOR kinase'. Curr. Biol. 15 (8): 702–13. doi:10.1016/j.cub.2005.02.053. PMID15854902.
- ^Karbowniczek M, Cash T, Cheung M, Robertson GP, Astrinidis A, Henske EP (July 2004). 'Regulation of B-Raf kinase activity by tuberin and Rheb is mammalian target of rapamycin (mTOR)-independent'. J. Biol. Chem. 279 (29): 29930–7. doi:10.1074/jbc.M402591200. PMID15150271.
- ^Yee WM, Worley PF (February 1997). 'Rheb interacts with Raf-1 kinase and may function to integrate growth factor- and protein kinase A-dependent signals'. Mol. Cell. Biol. 17 (2): 921–33. doi:10.1128/mcb.17.2.921. PMC231818. PMID9001246.
- ^Movilla N, Crespo P, Bustelo XR (October 1999). 'Signal transduction elements of TC21, an oncogenic member of the R-Ras subfamily of GTP-binding proteins'. Oncogene. 18 (43): 5860–9. doi:10.1038/sj.onc.1202968. PMID10557073.
- ^ abWang S, Ghosh RN, Chellappan SP (December 1998). 'Raf-1 physically interacts with Rb and regulates its function: a link between mitogenic signaling and cell cycle regulation'. Mol. Cell. Biol. 18 (12): 7487–98. doi:10.1128/mcb.18.12.7487. PMC109329. PMID9819434.
- ^Ayroldi E, Zollo O, Macchiarulo A, Di Marco B, Marchetti C, Riccardi C (November 2002). 'Glucocorticoid-induced leucine zipper inhibits the Raf-extracellular signal-regulated kinase pathway by binding to Raf-1'. Mol. Cell. Biol. 22 (22): 7929–41. doi:10.1128/mcb.22.22.7929-7941.2002. PMC134721. PMID12391160.
- ^Truong AB, Masters SC, Yang H, Fu H (November 2002). 'Role of the 14-3-3 C-terminal loop in ligand interaction'. Proteins. 49 (3): 321–5. doi:10.1002/prot.10210. PMID12360521.
- ^Yuryev A, Ono M, Goff SA, Macaluso F, Wennogle LP (July 2000). 'Isoform-specific localization of A-RAF in mitochondria'. Mol. Cell. Biol. 20 (13): 4870–8. doi:10.1128/mcb.20.13.4870-4878.2000. PMC85938. PMID10848612.
- ^ abcVincenz C, Dixit VM (August 1996). '14-3-3 proteins associate with A20 in an isoform-specific manner and function both as chaperone and adapter molecules'. J. Biol. Chem. 271 (33): 20029–34. doi:10.1074/jbc.271.33.20029. PMID8702721.
- ^ abConklin DS, Galaktionov K, Beach D (August 1995). '14-3-3 proteins associate with cdc25 phosphatases'. Proc. Natl. Acad. Sci. U.S.A. 92 (17): 7892–6. Bibcode:1995PNAS..92.7892C. doi:10.1073/pnas.92.17.7892. PMC41252. PMID7644510.
- ^ abEwing RM, Chu P, Elisma F, Li H, Taylor P, Climie S, McBroom-Cerajewski L, Robinson MD, O'Connor L, Li M, Taylor R, Dharsee M, Ho Y, Heilbut A, Moore L, Zhang S, Ornatsky O, Bukhman YV, Ethier M, Sheng Y, Vasilescu J, Abu-Farha M, Lambert JP, Duewel HS, Stewart II, Kuehl B, Hogue K, Colwill K, Gladwish K, Muskat B, Kinach R, Adams SL, Moran MF, Morin GB, Topaloglou T, Figeys D (2007). 'Large-scale mapping of human protein-protein interactions by mass spectrometry'. Mol. Syst. Biol. 3 (1): 89. doi:10.1038/msb4100134. PMC1847948. PMID17353931.
- ^Autieri MV, Carbone CJ (July 1999). '14-3-3Gamma interacts with and is phosphorylated by multiple protein kinase C isoforms in PDGF-stimulated human vascular smooth muscle cells'. DNA Cell Biol. 18 (7): 555–64. doi:10.1089/104454999315105. PMID10433554.
- ^Ichimura T, Wakamiya-Tsuruta A, Itagaki C, Taoka M, Hayano T, Natsume T, Isobe T (April 2002). 'Phosphorylation-dependent interaction of kinesin light chain 2 and the 14-3-3 protein'. Biochemistry. 41 (17): 5566–72. doi:10.1021/bi015946f. PMID11969417.
- ^Liu YC, Elly C, Yoshida H, Bonnefoy-Berard N, Altman A (June 1996). 'Activation-modulated association of 14-3-3 proteins with Cbl in T cells'. J. Biol. Chem. 271 (24): 14591–5. doi:10.1074/jbc.271.24.14591. PMID8663231.
- ^Clark GJ, Drugan JK, Rossman KL, Carpenter JW, Rogers-Graham K, Fu H, Der CJ, Campbell SL (August 1997). '14-3-3 zeta negatively regulates raf-1 activity by interactions with the Raf-1 cysteine-rich domain'. J. Biol. Chem. 272 (34): 20990–3. doi:10.1074/jbc.272.34.20990. PMID9261098.
- ^Tzivion G, Luo ZJ, Avruch J (September 2000). 'Calyculin A-induced vimentin phosphorylation sequesters 14-3-3 and displaces other 14-3-3 partners in vivo'. J. Biol. Chem. 275 (38): 29772–8. doi:10.1074/jbc.M001207200. PMID10887173.
- ^Koyama S, Williams LT, Kikuchi A (July 1995). 'Characterization of the interaction of Raf-1 with ras p21 or 14-3-3 protein in intact cells'. FEBS Lett. 368 (2): 321–5. doi:10.1016/0014-5793(95)00686-4. PMID7628630.
- ^Chow CW, Davis RJ (January 2000). 'Integration of calcium and cyclic AMP signaling pathways by 14-3-3'. Mol. Cell. Biol. 20 (2): 702–12. doi:10.1128/MCB.20.2.702-712.2000. PMC85175. PMID10611249.
Further reading[edit]
- Reed JC, Zha H, Aime-Sempe C, Takayama S, Wang HG (1997). 'Structure-function analysis of Bcl-2 family proteins. Regulators of programmed cell death'. Adv. Exp. Med. Biol. 406: 99–112. doi:10.1007/978-1-4899-0274-0_10. PMID8910675.
- Geyer M, Fackler OT, Peterlin BM (2001). 'Structure–function relationships in HIV-1 Nef'. EMBO Rep. 2 (7): 580–5. doi:10.1093/embo-reports/kve141. PMC1083955. PMID11463741.
- Dhillon AS, Kolch W (2002). 'Untying the regulation of the Raf-1 kinase'. Arch. Biochem. Biophys. 404 (1): 3–9. doi:10.1016/S0003-9861(02)00244-8. PMID12127063.
- Greenway AL, Holloway G, McPhee DA, Ellis P, Cornall A, Lidman M (2004). 'HIV-1 Nef control of cell signalling molecules: multiple strategies to promote virus replication'. J. Biosci. 28 (3): 323–35. doi:10.1007/BF02970151. PMID12734410.
- Chen H, Kunnimalaiyaan M, Van Gompel JJ (2006). 'Medullary thyroid cancer: the functions of raf-1 and human achaete-scute homologue-1'. Thyroid. 15 (6): 511–21. doi:10.1089/thy.2005.15.511. PMID16029117.
External links[edit]
- Domain structure diagrams for Raf-1, A-Raf and B-Raf.
- c-raf+Proteins at the US National Library of Medicine Medical Subject Headings (MeSH)
- Human RAF1 genome location and RAF1 gene details page in the UCSC Genome Browser.
Retrieved from 'https://en.wikipedia.org/w/index.php?title=C-Raf&oldid=882660370'
doi: 10.1177/1947601911407323
PMID: 21779496
Monographs Editor: Eugenio Santos
This article has been cited by other articles in PMC.
Abstract
First identified in the early 1980s as retroviral oncogenes, the Raf proteins have been the objects of intense research. The discoveries 10 years later that the Raf family members (Raf-1, B-Raf, and A-Raf) are bona fide Ras effectors and upstream activators of the ubiquitous ERK pathway increased the interest in these proteins primarily because of the central role that this cascade plays in cancer development. The important role of Raf in cancer was corroborated in 2002 with the discovery of B-Raf genetic mutations in a large number of tumors. This led to intensified drug development efforts to target Raf signaling in cancer. This work yielded not only recent clinical successes but also surprising insights into the regulation of Raf proteins by homodimerization and heterodimerization. Surprising insights also came from the hunt for new Raf targets. Although MEK remains the only widely accepted Raf substrate, new kinase-independent roles for Raf proteins have emerged. These include the regulation of apoptosis by suppressing the activity of the proapoptotic kinases, ASK1 and MST2, and the regulation of cell motility and differentiation by controlling the activity of Rok-α. In this review, we discuss the regulation of Raf proteins and their role in cancer, with special focus on the interacting proteins that modulate Raf signaling. We also describe the new pathways controlled by Raf proteins and summarize the successes and failures in the development of efficient anticancer therapies targeting Raf. Finally, we also argue for the necessity of more systemic approaches to obtain a better understanding of how the Ras-Raf signaling network generates biological specificity.
Keywords: Raf kinases, signal transduction, cancer, apoptosis, kinase inhibitors
Introduction
The first raf (rapidly accelerated fibrosarcoma) gene was described in 1983 as a retroviral oncogene, v-raf, transduced by the murine sarcoma virus isolate 3611. A year later, an avian homolog, v-mil, was found in the MH2 retrovirus. These 2 transforming retroviruses encoded the first oncogene to be discovered with serine/threonine kinase activity. After the cellular proto-oncogene homologs, c-raf and c-mil, had been cloned, studies focused on elucidating the function of Raf proteins. They showed that c-Raf (also known as Raf-1) plays a critical role in mediating the cellular effects of growth factor signals.- Later on, Raf proteins were identified as direct activators of MEK, and effectors of Ras.- Thus, Raf proteins were placed as essential connectors between Ras and the MEK-ERK pathway (Fig. 1). Most subsequent work focused on understanding this role and the regulation of Raf proteins in detail, until new functions of Raf-1 in the regulation of apoptosis- and cell migration emerged in the last decade.
The prototypical Ras-Raf-MEK-ERK pathway. Activated receptor tyrosine kinases (RTKs) recruit the guanine nucleotide exchange factor SOS, which activates Ras proteins by exchanging GDP for GTP. Activated GTP-loaded Ras binds to Raf, initiating Raf activation. Active Raf phosphorylates and activates MEK, which in turn phosphorylates and activates ERK. While the phosphorylation cascade comprising Raf, MEK, and ERK is linear, ERK features more than 150 substrates both in the cytosol and nucleus. Protein interactions and phosphorylation reactions are modulated by a number of scaffolding proteins (see Fig. 5).
Three different Raf isoforms originating from 3 independent genes can be distinguished in mammals, Raf-1/c-Raf, B-Raf, and A-Raf. Raf-1 was the first isoform to be identified and for 20 years was the principal focus of attention on the proteins of the family. After the discovery 8 years ago of B-Raf mutations in different types of tumors, B-Raf moved into the limelight, resulting in a rapid increase of our knowledge of the biological functions of this isoform. On the other hand, still very little is known about A-Raf, and although it seems to share many of the properties of the other isoforms, its biological functions remain a mystery. All Raf proteins share MEK1/2 kinases as substrates. MEK1/2 in turn activate ERK1/2, and this pathway regulates many cellular functions such as cell proliferation, differentiation, migration, or apoptosis (for extensive reviews, see Wellbrock et al., Leicht et al., and Dhillon et al.).
In recent years, it has become clear that the initial view of the ERK pathway as a linear pathway is not accurate but that there are many different proteins interacting with the proteins of the pathway. These proteins regulate the pathway by mediating the crosstalk with other signaling pathways and the regulation of positive and negative feedback mechanisms. In this review, we focus on the mechanisms of Raf family regulation and the biological roles of Raf family kinases especially in cancer and with relation to Ras signaling. We also explore the role of the Raf proteins in the context of the coordinated signaling networks that ultimately are responsible for cellular responses both in normal and tumor cells. We end on how systems biology can help us to integrate the information gathered from the many years of research in the Ras-Raf pathway and how it can be used to address open questions.
Structure and Regulation of Raf Isozymes
Raf Structure
There are no Raf kinases in yeasts, and the phylogenetic oldest isoform is B-Raf, which appears in invertebrates. Mammals possess 3 Raf isoforms (Raf-1, B-Raf, and A-Raf), which share a common modular structure consisting of 3 conserved regions (CR) with distinct functions (Fig. 2). CR1 contains a Ras-binding domain (RBD), which is necessary for the interaction with Ras and with membrane phospholipids required for membrane recruitment, and a cysteine-rich domain (CRD), which is a secondary Ras-binding site and also necessary for the interaction of CR1 with the kinase domain for Raf autoinhibition. CR2 contains important inhibitory phosphorylation sites participating in the negative regulation of Ras binding and Raf activation. CR3 features the kinase domain, including the activation segment, whose phosphorylation is crucial for kinase activation. Unfortunately, the tertiary structure of a Raf holoenzyme has remained elusive, although the structures of the RBD and extended CR1 domains of Raf-1- and the CR3 domain of B-Raf and Raf-1 were solved. Functionally, the Raf structure can be split into a regulatory N-terminal region, containing the RBD, which is critical for activation as well as inhibitory phosphorylation sites, and a catalytic C-terminal region, which includes phosphorylation sites necessary for the kinase activation. The regulatory domain restrains the activity of the kinase domain,, and its removal results in constitutive oncogenic activation. However, the activity of the isolated Raf-1 kinase domain is subjected to further regulation and can be stimulated by phorbol esters, v-Src, and phosphorylation., This observation is in keeping with the finding that the most common oncogenic mutation in B-Raf, V600E, activates B-Raf kinase activity by mimicking phosphorylation of the activation loop that releases its inhibitory interaction with the ATP-binding domain.
Structure and regulatory phosphorylation sites of Raf proteins. (A) Common structure of the Raf proteins. Color-coded regions are described in the text. (B) Comparison of the structure and phosphorylation residues of the 3 Raf isoforms. Red residues indicate activating phosphorylation sites, black are inhibitory sites, and blue are sites that have been described as both activating and inhibitory. The major in vitro autophosphorylation sites in Raf-1 and B-Raf are in green.
The Raf-1 Activation Cycle
In the inactive state, Raf-1 is thought to exist in a closed conformation in which the N-terminal regulatory region folds over and occludes the catalytic region. This conformation is stabilized by a 14-3-3 dimer binding to an N-terminal site, phospho-S259 (pS259), and a C-terminal site, pS621. Although the activation process of Raf-1 is not completely understood, we can assume the following sequence of events (Fig. 3).
The Raf-1 activation/deactivation cycle. This scheme shows the salient steps in Raf-1 activation/deactivation. Activating events are coded red, inactivating processes are in black, and activation states are in blue. In quiescent cells, Raf-1 is phosphorylated on both 14-3-3 binding sites pS259 and pS621, and 14-3-3 maintains the closed inactive conformation. Upon membrane recruitment by activated Ras, pS259 is dephosphorylated by the corecruited phosphatases PP1 or PP2A. Subsequently, phosphorylation of the N-region and activation loop and homodimerization or heterodimerization (with B-Raf) cause full activation of Raf-1. Deactivation is initiated by pS338 inducing PP5 binding and dephosphorylation of pS338. In addition, ERK-mediated feedback phosphorylation suppresses Raf-1 catalytic activity. Eventually, PP2A (and maybe other unknown phosphatases) dephosphorylates the remainder of activating sites and the ERK feedback sites. Rephosphorylation of S259 allows intramolecular bidentate 14-3-3 rebinding and return to the inactive state.
- Dephosphorylation of pS259 at the cell membrane by specific phosphatases (PP2A, PP1) releases 14-3-3 from its N-terminal binding site in Raf-1, thereby allowing conformational changes to occur that unmask the RBD and CRD domains in the CR1 region to enable Ras binding and membrane recruitment.-
- Ras binding itself has several intricate facets. The RBD is essential for the selective binding to activated RasGTP, and this binding interface was extensively characterized structurally, and functionally. A notable observation was that in the absence of feedback, the Ras–Raf-1 association rate kinetics rather than the total affinity determined the extent of downstream ERK activation. However, negative feedback operating from ERK back to Ras activation negated the subtlety of the kinetic effects and rendered ERK activation transient. This negative feedback is mediated by ERK and its downstream substrate RSK terminating Ras activation by phosphorylating and inhibiting the guanine exchange factor SOS., In this context, it is interesting that the binding of Raf-1 to Ras can be accelerated by the scaffolding protein Sur-8/SHOC2. A complex between Sur-8/SHOC2 and the catalytic subunit of PP1c is an effector of the Ras family protein M-Ras, which can dephosphorylate the inhibitory pS259 in Raf-1 at the membrane. Thus, Ras proteins can cooperate in recruiting Raf-1 and specific activators. The CRD exhibits constitutive low affinity for Ras that does not discriminate between Ras activation states. The CRD is not sufficient but necessary for stable membrane recruitment and activation of Raf-1., These results suggest that the CRD may stabilize the primary recruitment of Raf-1 exerted by the RBD through forging interactions with the farnesyl lipid tails of Ras proteins., In addition, Raf translocation to the membrane is aided by the ability of all 3 Raf isoforms to interact with lipids.- In fact, as phosphatidic acid (PA) can bind to both Raf-1 and SOS, PA was suggested to nucleate Ras nanocluster formation in response to EGF. Furthermore, Ras isoforms reside in different microcompartments, which can influence interactions with Raf kinases., This spatial organization may profoundly influence the mechanism and kinetics of Raf activation by different Ras isoforms. Ras itself seems to be organized in short-lived, highly dynamic nanoclusters, which activate Raf-1 in a digital way; that is, each Ras nanocluster produces a constant output of Raf activity. However, as the number of Ras nanoclusters increases proportionally with growth factor concentrations, the overall output is analog. This conversion of an analog input signal into an analog output by means of a digital intermediate increases the fidelity of signal transmission across the cell membrane. Interestingly, Ras isoforms also can reside at and signal from endomembranes including the Golgi apparatus and endoplasmic reticulum., The subcellular localization modulates the efficiency with which different effector pathways are engaged but, in the case of Raf-1, also has a major influence on the dynamics of signaling. While increasing doses of EGF induced ERK activation in a linear fashion when signaling from the plasma membrane early after EGF treatment, the dose response became nonlinear and sigmoid when activated from the Golgi at later time points. Thus, according to this model, different EGF concentrations are translated into ERK activity in a highly linear, quantitative fashion at the cell membrane in the early phase of signaling. At later time points, when ERK is activated from the Golgi, the dose response curve becomes nonlinear with a less accurate input-output relationship. The physiological role of these dynamic differences was demonstrated in T cell selection, where strong ERK activation at the plasma membrane drove negative selection, whereas delayed signaling from the Golgi promoted positive selection.
- Phosphorylation of the activation segment in CR3 and the “N-region” (negative charge regulatory region) is upstream of CR3. The N-region contains the SSYY phosphorylation sites, which are not only essential for full kinase activation but also for interaction with the substrate MEK.- The kinases, which can phosphorylate Y341, include Src, and JAK family kinases. Mutation of Y341 severely compromises Raf-1 kinase activity, but detecting phosphorylation of this residue in response to physiological stimuli is difficult, and most studies used overexpressed or mutated tyrosine kinases. This observation may indicate that our current detection methods are insufficient or that the modification is instable or actually a different chemical entity that provides a negative charge such as a sulfate group. Phosphorylation of S338 is detectable reliably and used routinely as a surrogate marker for Raf-1 activation. Pak family kinases were reported to phosphorylate S338 in response to growth factor stimulation, and integrin activation. However, the role of Pak in Raf-1 activation was questioned because stimuli that activate Raf-1 did not necessarily activate Pak and because Pak phosphorylation did not automatically activate Raf-1. More recent work suggested that S338 could be an autophosphorylation site induced by dimerization or be a target for casein kinase 2 (CK2) recruited to Raf-1 and B-Raf by the scaffold KSR. Given that Raf kinases can be activated by many diverse stimuli, it is not surprising that key phosphorylation sites are targeted by several different kinases. Interestingly, S338 phosphorylation is not required for Raf-1 activation at the Golgi, suggesting that the activation mechanism at the Golgi membrane may be different from the activation mechanism at the plasma membrane. S338 phosphorylation itself only slightly elevates Raf-1 kinase activity and mainly seems to serve as a priming event that initiates further activating modifications. Both Ras and Raf-1 activation at the Golgi is delayed relative to the plasma membrane, indicating that the initial Raf-1 priming events may be different between these compartments. Alternatively, recent evidence suggests that H-Ras is activated at the plasma membrane and endoplasmic reticulum and then delivered to the Golgi. In this scenario, Raf-1 could be activated at the plasma membrane and travel to the Golgi bound to activated H-Ras. As pS338 rapidly recruits the protein phosphatase PP5 to Raf-1, this residue may become dephosphorylated during this journey. However, activity may be maintained, as in this state, Raf-1 already would have undergone dimerization with B-Raf or KSR, which can activate Raf-1 allosterically.- At the Golgi, Raf-1 and B-Raf can associate with the Raf kinase trapping to the Golgi (RKTG) protein, which inhibits Raf signaling by interfering with Raf binding to Ras and MEK., The existence of such Golgi-specific Raf regulatory proteins suggests that the Golgi may have developed its own means to regulate Raf activity. Finally, the phosphorylation of 2 sites in the activation loop is required for full activation, and activation by Raf heterodimerization, but the identity of the respective kinases is unknown. In addition, S508 in the Raf-1 activation loop is involved in MEK binding.
- Raf homodimerization and heterodimerization recently emerged as important regulatory mechanisms to drastically enhance the kinase activity and signaling of Raf. It is not entirely clear at which step in the activation dimerization occurs. Although it is part of the physiological activation mechanism, it may also provide an alternative route of Raf activation independent of N-terminal phosphorylation. Because of the mechanistic complexity and relevance for mutant Raf signaling and drug responsiveness, it is discussed in detail below.
- Deactivation is initiated by specific binding of PP5 to activated Raf-1, which results in the dephosphorylation of pS338, rephosphorylation of S259, and return into the inactive state. The phosphorylated N-region also serves as a binding site for the Raf kinase inhibitor protein (RKIP), which dissociates Raf-1 from its substrate MEK., In addition, Raf-1 is subjected to direct feedback phosphorylation by ERK on 6 sites, which inhibits the activation of Raf-1 by Ras and promotes the subsequent dephosphorylation of Raf-1 by PP2A and the return to the inactive state. A negative feedback from ERK to Raf-1 was also confirmed by a systematic analysis of feedback regulation of the ERK pathway based on mathematical modeling. However, ERK feedback phosphorylation also was described as stimulating Raf-1 activity. The reason for this contradiction is not clear. While Dougherty et al. reported 6 feedback sites, Balan et al. identified a subset of 3 of the 6 sites. Two of these 3 sites were also identified as stimulatory phosphorylation sites in A-Raf, raising the interesting possibility that the phosphorylation of a subset of ERK feedback sites has a positive effect, whereas phosphorylation of the full complement is inhibitory. Such a mechanism could be a simple way to dynamically regulate strength and duration of ERK signaling, where early incomplete phosphorylation would boost Raf-1 activity, while later complete phosphorylation would switch Raf-1 activation off. Another important negative regulation of Raf-1 is phosphorylation by cyclic AMP–activated kinase (PKA). This topic was recently reviewed and therefore is only presented briefly. Raf-1 is a direct PKA substrate, and different studies found several sites in which PKA can phosphorylate Raf-1. The phosphorylation of S43 interferes with binding to Ras, while phosphorylation of S233 and S259 enhances the binding of 14-3-3 and suppresses catalytic activity., The phosphorylation of S621 has a dual role. It decreases Raf-1 kinase activity, but its inhibitory function is converted into an essential component of Raf-1 activity by 14-3-3 binding. S259 also was reported to be phosphorylated by Akt, but this observation could not be reproduced.,
A-Raf Regulation
A-Raf is generally thought to be regulated similarly to Raf-1, but important differences have emerged. A-Raf is only weakly activated by oncogenic H-Ras and Src and also displays low kinase activity towards MEK. The reason for the lower responsiveness to H-Ras is the exchange of an arginine for a lysine at position 22 in the A-Raf RBD, which weakens the binding of A-Raf to H-Ras. In addition, the low kinase activity may be unique nonconserved amino acid residues in the N-region. Mutation of Y296 in the N-region led to a constitutively active kinase, and molecular modeling showed that Y296 promotes a tighter interaction between the N-region and the catalytic domain, which may stabilize the closed conformation. Subsequently, a systematic phosphorylation site analysis revealed several interesting findings: S432 located between the ATP-binding domain and activation loop was found critical for MEK binding and A-Raf signaling. Surprisingly, activation loop phosphorylation did not contribute to mitogen-induced activation. Finally, a cluster of phosphorylation sites between amino acids 248 and 267, which stimulated activation, facilitated A-Raf dissociation from the plasma membrane. These findings raise some intriguing aspects. First, activation loop phosphorylation, which is a widespread mechanism in the catalytic activation of kinases, may be less critical in Raf regulation, as corresponding structural reorganizations may be caused by 14-3-3 proteins binding to the Raf kinase domain (see below). Second, the fact that activating phosphorylation events release A-Raf from the plasma membrane suggests that while initial activation may occur at the membrane, downstream signaling proceeds in other subcellular compartments. In this context, both A-Raf and Raf-1 have been found in different subcellular compartments including mitochondria, endosomes, and the Golgi apparatus. It is unknown whether A-Raf is regulated by PKA.
B-Raf Regulation
B-Raf activation appears much simpler. In fact, Ras and 14-3-3 binding are likely to be the only major requirement for B-Raf activation., The N-region is already negatively charged because of the presence of aspartate at the position corresponding to Raf-1’s YY340/1 (DD448/9) and the constitutive phosphorylation of S446 (corresponding to the S338 of Raf-1). Phosphorylated S446 neutralizes the inhibitory role of N-terminal domain towards the catalytic region and in conjunction with D449 allows the catalytic domain to adopt a stabilized 3-dimensional conformation. Phosphorylation of S365 (corresponding to S259 in Raf-1) impairs B-Raf activity, and its mutation to a nonphosphorylatable residue can even overcome the debilitating effects of charge-neutralizing mutations in the N-region., In contrast to Raf-1 S259, B-Raf S365 is unlikely to be phosphorylated in cells upon cAMP stimulation, but another site, S429, is a potential target for PKA. Importantly, B-Raf can be inhibited or activated by PKA depending on the levels of 14-3-3 expression, which need to be high for permitting activation. A second pathway may involve the PKA-mediated activation of Rap1, which was reported to bind and activate B-Raf., However, these results are disputed, as many physiological signals that induce Rap1 activity fail to activate B-Raf. Possible reasons for this discrepancy are unclear and are discussed below.
The Role of Ras Family Proteins in Raf Isoform Regulation
The common and key step in the activation of all 3 Raf isoforms is membrane recruitment by a Ras family protein. Membrane translocation triggers further activation events, such as the binding of PP2A to dephosphorylate the inhibitory pS259 site in Raf-1 (and presumably the corresponding sites in A-Raf and B-Raf) and the colocalization with the kinases responsible for the multiple activating phosphorylations as discussed above. The sequences forming the binding interface are well conserved in the Raf as well as Ras family. Hence, it is not surprising that several members of the Ras family can bind Raf kinases. A systematic comparison of the ability of different Ras family members to activate Raf isoforms showed that H-Ras, N-Ras, and K-Ras could stimulate all 3 Raf isoforms and were the only Ras proteins that could activate B-Raf. In contrast, A-Raf also could be activated by R-Ras3, while Raf-1 was the most promiscuous isoform, responding also weakly to R-Ras3, Rit, and TC21. In contrast, Rap1/2, Rin, and Rheb were ineffective. The ability to activate Raf generally corresponded with the binding affinity. Only H-Ras, N-Ras, and K-Ras led to a stimulation of the endogenous ERK pathway in HEK293 cells, while lower affinity interactions only could stimulate ERK when either Raf or ERK was overexpressed. These results suggest that cell type–specific expression stoichiometry of Ras isoforms and Raf-ERK pathway components potentially could generate a rich variety of ERK activation dynamics that could allow cells to respond to different growth factors with precisely tuned ERK activation. An example is Rheb, which was reported to interact with Raf-1 and B-Raf and to suppress their kinase activity by reducing N-region phosphorylation and heterodimerization., Interestingly, PKA-mediated phosphorylation of Raf-1 on S43 increases Raf-1 affinity for Rheb and thereby could contribute to the inhibitory effects of PKA by diverting Raf-1 from H-Ras to Rheb.
Although this hypothesis is conceptually appealing, the experimental evidence for specific engagement of Raf isoforms by different Ras family members is often controversial. A case in particular is Rap1, which was initially isolated as a repressor of K-Ras transformation. Rap1 binds to Raf-1, but the functional consequences are disputed. Constitutively active Rap1 can inhibit Raf-1 when overexpressed, while at normal expression levels and in response to physiological stimulation, Rap1 did not regulate Raf-1., In addition, Rap1 was reported to activate B-Raf, and mediate cAMP stimulation of ERK and neuronal differentiation of PC12 cells. However, this finding was not reproduced in other studies, and the issue remains open. A possible explanation for the differential effect of Rap1 and Ras on B-Raf and Raf-1 is the difference in affinity to the CRD domains. Rap1 has high affinity for the Raf-1 CRD domain and low affinity for the B-Raf CRD, whereas Ras has low affinity for both Raf-1 and B-Raf CRD domains. Swapping the CRD domains showed that the B-Raf CRD conveyed susceptibility to activation by Rap1, while the Raf-1 CRD abolished it. An alternative, but not mutually exclusive, mechanism could be the failure of Rap1 to target Raf-1 to the membrane compartment, where S338 phosphorylation can take place. Correct targeting needs Y341 phosphorylation or the negative charge that B-Raf carries at this position. Introducing a negatively charged residue at this position or redirecting Rap1 to lipid rafts could overcome the deficiency of Rap1 to activate Raf-1. A recent study using statistical modeling based on Bayesian inference revealed a role for Rap1 in EGF-stimulated ERK activation by supporting Raf-1 and B-Raf heterodimerization. The exact mechanism needs to be elucidated. Apart from direct Rap1 binding to B-Raf, it also could involve indirect effects such as a recruitment of a common scaffold by Rap1. In natural killer lymphocytes, Rap1 was recently shown to bind to the scaffolding protein IQ motif containing GTPase-activating protein 1 (IQGAP1) and to assemble a signaling complex that activated B-Raf, Raf-1, and ERK. Thus, the contradictory findings on Rap1 regulation of Raf proteins may eventually be reconciled when considering indirect mechanisms such as the involvement of scaffolds.
Regulation of Raf Isoform Expression by Differential Splicing
All 3 Raf isoforms are regulated at the level of protein expression. Alternative splicing gives rise to multiple B-Raf isoforms differentially expressed in various tissues., B-Raf activity is also regulated by splicing. B-Raf isoforms containing exon 8b are more phosphorylated on the inhibitory S365 site, leading to an increased interaction with 14-3-3 and strengthening the inhibitory interaction between N-terminal regulatory domain and kinase domain, altogether resulting in lower kinase activity. With respect to A-Raf, the 2 splice isoforms described so far, DA-Raf1 and D-Raf2, lack the kinase domain and act as dominant inhibitory mutants of Ras and ARF GTPases. As a consequence, DA-Raf1 is a positive regulator of myogenic differentiation by mediating the inhibition of the ERK pathway required for differentiation. Raf-1 also has a known splice variant preferentially expressed in the muscle and brain.
Raf Regulation by 14-3-3 Proteins
Key regulatory phosphorylation sites in Raf kinases are also binding sites for the scaffolding protein 14-3-3. 14-3-3 is an obligatory dimer forming a rigid half-barrel structure that interacts with other proteins in a phosphorylation-dependent, bidentate way, constraining the conformation of the binding partner. In Raf kinases, 14-3-3 can stabilize both the inactive and the activated state. This dual property confounds the analysis of 14-3-3 effects on Raf kinase regulation and can explain many of the controversies in the literature.
In the inactive configuration, 14-3-3 binds to conserved phosphorylation sites in the N- and C-terminus of the Raf kinase domain (pS259 and pS621 in Raf-1; pS365 and p729 in B-Raf; pS214 and pS576 in A-Raf).,- This bidentate interaction is thought to physically clamp the regulatory domain to the kinase domain. Thus, it is not surprising that the 14-3-3 binding sites are targeted by kinases that inhibit Raf activation. Both protein kinase A (PKA) and B (PKB/Akt) were reported to induce phosphorylation of the N-terminal 14-3-3 binding site of Raf-1 and B-Raf.,-,- However, the relevance of Akt phosphorylation remains disputed, as Akt does not phosphorylate S259 in most physiological scenarios., While PKA is a bona fide S259 kinase, it is not responsible for the constitutive phosphorylation of S259, and the kinase that maintains basal S259 phosphorylation in cells is still unknown. The C-terminal 14-3-3 binding residue S621 is targeted by PKA, AMP-activated protein kinase (AMPK), autophosphorylation,, and probably other yet unidentified kinases. The exact role of S621 is still not clear. It was reported to inhibit Raf-1 kinase but also to be essential for kinase activity, and for the stability of the Raf-1 protein. Recent data show that mutations that preclude 14-3-3 binding to S621 inactivate Raf-1 by specifically disrupting its capacity to bind to ATP rather than by gross conformational alteration, as indicated by the intact ability to bind MEK. Phosphorylation of S621 inhibits Raf-1 catalytic activity in vitro, but addition of 14-3-3 proteins completely reverses this inhibition. 14-3-3 binding requires the phosphorylation of S621, and this interaction is essential for Raf-1 kinase activity, but S621 phosphorylation in the absence of 14-3-3 does not support kinase activity. These data explain the dual role of S621 phosphorylation and suggest that 14-3-3 may serve as a switch that can convert an inactive Raf-1 population phosphorylated on S621 into a kinase-competent state. Although B-Raf catalytic activity is less dependent on 14-3-3 binding, its biological activity is dependent on 14-3-3 binding,, and 14-3-3 binding can switch inhibitory phosphorylation of B-Raf by PKA into activation. An additional role of 14-3-3 in Raf kinase activation is related to its enhancement of Raf-1 homodimerization and Raf-1/B-Raf heterodimerization, which elevates Raf-1 kinase activity, and is discussed below.
Raf Homodimers and Heterodimers
Dimerization is a common motif in the activation of kinases. Homodimerization was initially highlighted as a potentially important step of Raf-1 activation by 2 studies showing that a forced interaction of Raf-1 monomers tagged with inducible dimerizing tags robustly induced kinase activity., Both studies proposed that active Ras would promote the formation of dimers. This hypothesis was later extended to heterodimerization between Raf-1 and B-Raf, which was found to be inducible by active Ras. As mutation of S621 abrogated Raf heterodimerization, the authors speculated that 14-3-3 binding to pS621 was necessary for Raf heterodimerization. Although these initial studies showed that homodimerization and heterodimerization can activate Raf kinases, they failed to show that the interaction can take place between endogenous proteins when stimulated by physiological mitogens. This was achieved by Rushworth et al., who demonstrated that endogenous B-Raf and Raf-1 heterodimerize in multiple cell lines in response to mitogens. Biochemical fractionation of Raf heterodimers from homodimers and monomers showed that Raf-1–B-Raf heterodimers accounted for the majority of the mitogen-induced kinase activity. Remarkably, the heterodimers represented less than 1% of the total B-Raf pool but exhibited approximately 30-fold elevated kinase activity. Heterodimerization was enhanced by 14-3-3, but not a dimerization-negative 14-3-3 mutant, suggesting that the 14-3-3 dimer crosslinks Raf-1 and B-Raf by binding to the C-terminal sites on each kinase. This observation suggests a mechanism for how 14-3-3 can stabilize both inactive and active Raf-1 conformations. In the inactive conformation, 14-3-3 clasps the Raf-1 regulatory to the kinase domain via intramolecular binding to pS259 in the N-terminus and p621 in the C-terminus. Binding to activated Ras displaces 14-3-3 from pS259, leaving one 14-3-3 arm free to contact a 14-3-3 binding site in B-Raf to facilitate heterodimerization. In addition, Raf heterodimerization is also regulated by ERK-mediated feedback phosphorylation of B-Raf., The feedback phosphorylation mainly serves to limit the lifetime of B-Raf–Raf-1 heterodimers, and mutation of the relevant sites enhances ERK signaling and the associated biological activities. Other regulators of Raf heterodimerization include KSR1 and MLK3, which both enhance heterodimerization. MLK3 was originally described as an activator of JNK, but by activating B-Raf, it may serve as an integrating hub between the ERK and JNK pathways.
Heterodimerization also may play a pathophysiological role in cancer. When B-Raf mutations were discovered in cancer, a puzzling observation was that while the most frequent mutation, V600E, massively stimulated B-Raf kinase activity, several less frequent mutations activated B-Raf only mildly or not at all. However, even the low-activity B-Raf mutants could hyperstimulate the ERK pathway. Intriguingly, this activation was dependent on the presence of Raf-1, suggesting that low-activity B-Raf mutants require Raf-1 to activate the ERK pathway. A subsequent study confirmed the transactivation hypothesis by demonstrating that the low-activity B-Raf mutants found in cancer indeed could activate Raf-1 merely by forming dimers. While physiological heterodimerization is induced by Ras activation,, oncogenic B-Raf mutants constitutively dimerized with Raf-1. The mechanism of activation by heterodimerization is incompletely understood. Mutational analysis showed that it does not depend on N-region phosphorylation but requires the activation loop phosphorylation sites. This finding is difficult to reconcile with the observation that Ras-mediated activation of Raf-1 requires N-region phosphorylation., The ability of Ras to induce Raf heterodimerization should overcome the requirement for N-region phosphorylation for Raf activation. Thus, more work will be needed to elucidate the mechanism of how heterodimerization activates Raf kinase activity. Regardless of the exact mechanism, dimerization clearly has biological effects. Increasing the lifetime of Raf heterodimers by mutating the T753 ERK feedback phosphorylation site in B-Raf augmented the ability of nerve growth factor (NGF) to induce neuronal differentiation of PC12 cells. Furthermore, the mutation of ERK feedback phosphorylation sites and the concomitant increase in Raf heterodimer levels enhanced the transforming potential of oncogenic B-Raf with intermediate or low kinase activity, which depend on Raf-1 for transformation, without affecting transformation by the high-activity B-Raf V600E mutant.
An important role for Raf heterodimers was recently described in response to Raf kinase inhibitors. The original observation that Raf kinase inhibitors paradoxically could hyperactivate the Raf pathway is now explainable by the ability of Raf kinase inhibitors to promote Raf heterodimerization and activation., These findings are discussed below.
Signaling Downstream of Raf
Raf-Catalyzed MEK Phosphorylation
Despite much effort to identify Raf substrates, so far, the only bona fide physiological substrates of Raf are MEK1 and MEK2. Activated Raf kinases phosphorylate both MEK isoforms on 2 residues in the activation loop (S217 and S221). Phosphorylation on these 2 sites increases MEK activity, which in turn can bind, phosphorylate, and activate ERK (Fig. 1). Although all family members have the ability to bind and phosphorylate MEK in vitro, the activities towards MEK differ widely. B-Raf has the strongest activity towards MEK, followed by Raf-1 and A-Raf, whose MEK kinase activity is barely detectable. B-Raf possesses higher basal activity partially because it is “primed” for activation due to the aforementioned constitutive phosphorylation of the S445 residue and the negatively charged amino acids at positions in the N-region that need to be phosphorylated in Raf-1 and A-Raf to achieve activation. Furthermore, of all Raf isoforms, B-Raf has the strongest binding affinity for MEK. MEK is present in B-Raf protein-protein complexes even in starved cells, representing a preassembled complex ready to be activated.
The interaction of MEK with Raf-1 is also regulated by scaffolding proteins, such as KSR (see below), and by phosphorylation. The Rac effector kinase PAK1 can phosphorylate MEK1 on S298, which enhances the interaction with Raf-1 as well as ERK2. Feedback phosphorylation of T292 by activated ERK prevents the phosphorylation of S298 and limits MEK activation by Raf-1. This modulation is likely part of the biochemical basis for the cooperation between Ras and Rac proteins in cell transformation., Structural studies revealed an interesting role for T292. T292 only occurs in MEK1, but it can also downregulate the activity of MEK2 when T292-phosphorylated MEK1 heterodimerizes with MEK2. The implications of this differential regulation are not yet fully understood but indicate that MEK1 and MEK2 are not equivalent in terms of their regulation, and in a wider context, that dimerization of kinases also can exert negative control.
An unresolved question is why the activities of the comparatively poor MEK kinases Raf-1 and A-Raf are regulated in a much more complicated manner than the activity of the superior MEK kinase B-Raf. One possibility is that Raf-1 and A-Raf have other, yet unidentified substrates that are the real targets of their elaborate regulation. A more subtle possibility is that Raf-1 and A-Raf act as modulators that fine tune the ability of B-Raf to activate the ERK pathway. The discovery of B-Raf–Raf-1 heterodimers and their unfolding biological roles supports the latter view.
MEK-Independent Raf Signaling
A wealth of experimental data suggests that MEK is the only bona fide Raf substrate and that B-Raf is the main MEK kinase in vivo. This assessment is corroborated by phylogenetic comparisons. The single Raf homologs in invertebrates (lin-45 in Caenorhabditis elegans and D-Raf in Drosophila) are much closer to B-Raf in terms of sequence, suggesting that B-Raf is the archetypal MEK kinase, whereas A-Raf and Raf-1 may have been evolved towards MEK-independent functions.,
Alternative Raf substrates
Although MEK is the only commonly accepted Raf substrate, several other potential Raf-1 substrates were described. The adenylyl cyclases (AC) type 6, 5, and 2 were proposed to be phosphorylated and activated by Raf-1 independent of MEK.- As ACs generate cAMP, which activates PKA, their stimulation would initiate a negative feedback to Raf-1. Another proposed Raf-1 substrate is the retinoblastoma tumor suppressor protein (Rb) (Fig. 4). The inactivation of Rb by cyclin-dependent kinases 4 and 6 marks the irreversible commitment of the cell to divide. Raf-1 was reported to directly phosphorylate and inactivate Rb, leading to cell cycle progression. Disruption of the Rb–Raf-1 complex by interfering peptides or small molecules suppressed the growth of experimental tumors and associated angiogenesis in nude mice. Two other potential Raf-1 substrates regulate myosin contractility. One is myosin phosphatase (MYPT), which binds to Raf-1 and is phosphorylated by Raf-1, leading to MYPT inhibition and enhanced cell motility. Raf-1 can phosphorylate MYPT on the same site as the Rho effector kinase Rok-α and myotonic dystrophy protein kinase (MDPK). Interestingly, both kinases are regulated by Raf-1; Rok-α via binding and MDPK were reported to be phosphorylated and activated by Raf-1 directly. Finally, Raf-1 was described to phosphorylate cardiac troponin T, which regulates the contractile function of cardiomyocytes.
New Raf-1 signaling pathways that depend on protein interactions but not Raf-1 kinase activity or MEK. Download videomix pro for android. Raf-1 can suppress apoptosis in a MEK-independent fashion in several ways: 1) by binding to and inhibiting ASK1; 2) by suppressing cytochrome C release through voltage-dependent anion channels (VDACs) at the mitochondria; 3) by acting as a scaffold to recruit PKCθ to phosphorylate and inactivate BAD; 4) by inhibiting the mammalian MST2 pathway; and 5) by inhibiting Rok-α–induced Fas maintenance and clustering at the cell membrane. In addition, the inhibition of Rok-α by Raf-1 is also required for motility by regulating the actin cytoskeleton and for skin tumorigenesis by preventing keratinocyte differentiation and sustaining Myc expression.
Lessons from raf gene knockout mice
However, while the functional role of these putative Raf-1 substrates is not yet clear, new Raf effector pathways were discovered in which Raf-1 kinase activity is dispensable and the regulation occurs through association (Fig. 4). These discoveries were facilitated by the availability of conventional and conditional raf knockout mice.a-raf knockout mice are born alive but show neurological and intestinal defects of different severity depending on the genetic background. In contrast, b-raf–deficient embryos die around midgestation because of vascular defects in the placenta. Epiblast-restricted ablation, which leaves b-raf intact in the placenta but knocks the gene out in the embryo, resulted in live born animals that within 3 weeks succumb to a violent neurodegenerative disease.raf-1–deficient embryos show increased apoptosis of embryonic tissues or, more selectively, of the fetal liver depending on the genetic background., These divergent phenotypes show that Raf-1, B-Raf, and A-Raf serve distinct essential functions in embryonic development. It subsequently became clear that a main function of Raf-1 was to restrict caspase activation in response to selected stimuli, notably Fas stimulation, pathogen-mediated macrophage apoptosis, and erythroid differentiation., The ERK pathway can antagonize apoptosis in a number of ways, including the expression of caspase inhibitors and the neutralization of proapoptotic Bcl-2 family members. A further prominent prosurvival molecule, the transcription factor NFκB, was proposed as a downstream target of Raf-1.- It is still unclear how Raf kinases activate NFκB, but it is likely through the induction of autocrine factors., Importantly, neither MEK/ERK nor NFκB activation is altered in raf-1– or a-raf–deficient cells and embryos, indicating that the prosurvival roles of Raf-1 and A-Raf do not depend on these functions. What are then the essential downstream targets of Raf in apoptosis?
Raf-1 regulates apoptosis through multiple targets: Fas, ASK1, MST2, and Rok-α
A mitochondrial pool of Raf-1 was shown to protect cells from apoptosis.- Raf-1 could be targeted to the mitochondria via interaction with Bcl-2 when Bcl-2 was overexpressed. In addition, selected growth factors were reported to promote Raf-1 translocation to the mitochondria via p21-activated kinase (PAK)–induced phosphorylation on S338., However, it still is unclear how mitochondrially localized Raf-1 could prevent apoptosis. Raf-1 facilitates the phosphorylation and inactivation of the proapoptotic Bcl-2 family member BAD, although this is likely an indirect effect mediated by ERK-activated RSK., In addition, Raf-1 serves as a scaffold to recruit protein kinase C theta (PKCθ) to phosphorylate BAD. Another described mechanism is the direct interaction between Raf-1 and mitochondrial voltage-dependent anion channels (VDACs), which may prevent the release of cytochrome C from the mitochondria.
Gene ablation experiments in mice demonstrated that Raf-1 is required for survival and protection against apoptosis., Interestingly, reconstituting Raf-1–/– mice with a mutated Raf-1 (Raf-1 YY340/1FF), which has no detectable kinase activity towards MEK, fully rescued the apoptotic phenotype and produced viable mice. Tracking the cause revealed several mechanisms, which may operate in a tissue-specific manner, but none of which requires Raf-1 kinase activity. One is the control of the Rho effector kinase Rok-α, which is hyperactivated and mislocalized to the membrane in Raf-1 knockout cells., Hyperactive Rok-α causes a defect in the internalization of the Fas death receptor, which maintains high levels of Fas in the plasma membrane, leading to increased Fas sensitivity. The other targets of Raf-1 in apoptosis suppression are 2 proapoptotic kinases, ASK1 and MST2, which are inhibited by Raf-1 through direct binding (Fig. 4). These inhibitions do not require Raf-1 kinase activity but are solely mediated by binding. ASK1 is a protein kinase that works upstream of JNK and p38 to promote apoptosis induced by stress or by death receptors, such as the TNF-α receptor or Fas., It was reported that in human endothelial cells, Raf-1 mediates the protective effect of basic fibroblast growth factor (bFGF) against doxorubicin-induced apoptosis by binding to and inhibiting ASK1 at the mitochondria. Mutation of S338/339 in the N-region abolished association and protection. The mechanism of inhibition is not known, but the pathophysiological relevance of ASK1 inhibition by Raf-1 was demonstrated in a mouse model of heart disease. Knocking out the Raf-1 gene specifically in the heart muscle resulted in ventricular dilation and fibrosis caused by an increase in cardiomyocyte apoptosis. These pathological changes could be prevented by also knocking out ASK1.
The other Raf-1–inhibited proapoptotic kinase, MST2, was identified in a proteomics screen of Raf-1–associated proteins. The mechanism of how Raf-1 regulates MST2 was elucidated. MST2 activation involves dimerization and autophosphorylation of the activation loop. Raf-1 binds to the SARAH domain of MST2, thereby interfering with dimerization, and also recruits a phosphatase that dephosphorylates MST2. Raf-1 kinase activity is not required, and kinase-dead Raf-1 mutants also can inhibit MST2 activation. Consequently, MST2 activity is constitutively elevated in Raf-1 knockout cells and hyperactivatable by Fas stimulation or expression of RASSF1A. RASSF1A can disrupt the Raf-1–MST2 complex and promote the assembly of a proapoptotic signaling complex consisting of RASSF1A, MST2, LATS1, and YAP. In this context, MST2 phosphorylates LATS1, which phosphorylates YAP, thereby enabling YAP to interact with p73. The YAP-p73 complex binds to the promoter and activates the expression of the proapoptotic BH3 gene PUMA, culminating in the induction of apoptosis. This pathway was mapped using proteomics to track protein interactions that change in response to proapoptotic signals. This study flagged important gaps in our understanding of MST2 signaling. First, it showed that the MST2 pathway in mammalian cells shares the core kinase module MST2-LATS but differs from the orthologous Hippo pathway described genetically in Drosophila melanogaster in regard to upstream regulators and downstream effectors. This finding was subsequently confirmed by other studies., Second, it revealed the double nature of YAP as an oncogene as well as tumor suppressor. While in the liver, YAP is a potent oncogene,, YAP also can suppress oncogenesis by stimulating apoptosis in response to DNA damage- or expression of RASSF1A. RASSF1A is a major tumor suppressor gene that is altered in the majority of human cancers usually by gene silencing due to promoter methylation and much less frequently by mutation., Thus, being regulated by a major tumor suppressor pathway and a major mitogenic pathway, MST2 may have a critical role in coordinating apoptotic and transforming signals. Therefore, it is not surprising that MST2 is also targeted by the phosphatidylinositol 3-kinase (PI3K)/Akt survival pathway. Akt activation is required to curtail MST2 activation during growth factor stimulation. Akt phosphorylation of MST2 stimulates the dissociation of MST2 from RASSF1A and rebinding of MST2 to Raf-1. The role of Raf-1 is reminiscent of the role of the Myc proto-oncogene, which can stimulate transformation and apoptosis. Ras binding to Raf-1 enables Raf-1 to activate the MEK-ERK pathway and promote proliferation but at the same time dissociates the MST2–Raf-1 complex and promotes apoptosis. Coupling cell proliferation to the risk of cell death seems paradoxical but makes perfect sense for a multicellular organism in which the unlicensed proliferation of cells can cause severe diseases including cancer. Interestingly, B-Raf fails to bind and regulate MST2. Therefore, MST2 regulation by Raf is absent in Drosophila melanogaster, whose single Raf gene is most closely related to B-Raf. These observations suggest that MEK kinase activity was the primary function of Raf and that the ability to inhibit MST2 was acquired later in evolution.
Raf-1 regulates cell motility and differentiation through Rok-α
Another defect observed when Raf-1 was knocked out specifically in keratinocytes was retarded wound healing and migration of keratinocytes. Again, this novel function of Raf-1 can also be carried out by a kinase-dead mutant, and just like prosurvival, it involves the inhibition of another kinase. The target of Raf-1 in motility is Rok-α.Raf-1 knockout fibroblasts and keratinocytes have a contracted appearance, have a defective cytoskeleton characterized by tight cortical actin bundles, and fail to migrate. Chemical inhibition of Rok-α or expression of a dominant-negative Rok-α mutant rescues all these defects of the Raf-1–deficient cells, indicating that Rok-α is the only target of Raf-1 in motility. Interestingly, investigating the mechanism of Rho-α inhibition by Raf-1 revealed a critical role for the Raf-1 CRD. Rok-α, like Raf-1, is regulated by autoinhibition, and its C-terminal regulatory region features a domain highly homologous to the CRD found in Raf-1. Indeed, dependent on an intact CRD, the Raf-1 regulatory domain could crossregulate Rok-α by binding to the Rok-α kinase domain and repressing its function. The biological relevance of this interaction was borne out in a Ras-induced skin tumor model in mice. In this model, the inhibition of Rok-α by Raf-1 was required for Ras transformation by maintaining the dedifferentiated state of the tumor cells.
A-Raf signaling independent of MEK
A-Raf is the family member with the poorest kinase activity towards MEK and hence is likely to have functions outside of the classic ERK pathway. A-Raf strongly interacts with and inhibits MST2, again independently of kinase activity. Interestingly, this inhibition is contingent on the splice factor hnRNP H maintaining the expression of a full-length A-Raf protein. hnRNP H is often overexpressed in tumors. Downregulation of hnRNP H results in alternative splicing of the a-raf transcript that abolishes the expression of full-length A-Raf protein. Both A-Raf and MST2 are localized at the mitochondria in tumor cell lines as well as primary tumors.
In a yeast 2-hybrid screen, pyruvate kinase M2 (PKM2) was identified as an A-Raf binding partner. PKM2 is an embryonic splice form of PKM that is aberrantly re-expressed in cancer and responsible for the aerobic glycolysis, also known as the Warburg effect, in cancer cells. A-Raf–mediated transformation increased the activity of PKM2 by promoting the transition of PKM2 from the low-activity dimeric to the highly active tetrameric form, and PKM2 enhanced A-Raf–induced cell transformation., These findings potentially link A-Raf to the regulation of energy metabolism and cell transformation, a topic that is increasingly recognized as critical for tumorigenesis.
Scaffolds and Modulators of Raf Signaling
For a long time, the Ras/ERK signaling pathway was depicted as a linear pipeline. Over the years, it became clear that signaling pathways form networks consisting of multiprotein nodes at various subcellular compartments., A main question is how do these networks generate biological specificity? Much of this coordination depends on controlling protein-protein interactions by scaffold proteins that regulate the intensity, amplitude, and spatial specificity of the ERK pathway signal., Scaffolds act as docking platforms and anchors of the signaling components, bringing together the different modules of the cascade. Thus, by facilitating interactions between their clients, they decrease reaction rates to the first order, and they also reduce the number of tiers of the cascade, causing the input/output responses to become more linear. They insulate the clients from other pathways but also can connect pathways by binding components of different pathways. They can target their clients to different localizations, thereby increasing the variety of signals regulated by the cascade. There is now experimental evidence that scaffolds can link different localizations of Ras activation with the phosphorylation of specific ERK substrates., Feedback phosphorylation of the EGF receptor (EGFR) by ERK involved the IQ motif containing GTPase-activating protein (IQGAP) scaffold, while the phosphorylation of cytosolic phospholipase A2 (cPLA2) utilized KSR1 or Sef-1 when ERK was activated by Ras localized at the plasma membrane or Golgi, respectively. In addition, scaffolds seem to preferentially bind dimerized ERK and direct ERK to cytosolic substrates, whereas ERK dimerization is not required for the phosphorylation of nuclear substrates. The requirement for ERK dimerization is likely related to the overlap between the binding sites for substrates and scaffolds, implying that a dimer is necessary to simultaneously engage the scaffold and substrate.
Therefore, scaffolds can have a huge impact on the biochemical and biological behavior of the ERK pathway., However, our knowledge of their role in the functional modulation of the pathway and their exact mechanism of action is still limited. One problem is that scaffolds are quite difficult to study experimentally. Their function is highly dependent on concentrations and the stoichiometric ratios with respect to their client proteins, and both downregulation and overexpression have similar effects, as both conditions reduce the number of functional complexes. Scaffold proteins of the ERK pathway were extensively reviewed.,- We, therefore, only describe selected examples that allow us to outline salient functions of scaffolding proteins in the regulation of the ERK pathway (Fig. 5). For convenience, we have classified them by their major known functions. However, occasionally, these functions overlap and will expand as more details become known.
Scaffolding proteins in Raf-MEK-ERK signaling. Scaffolding proteins form Raf-MEK-ERK signaling platforms at different subcellular localizations. See text for details.
Scaffolds as Regulators of ERK Pathway Activity
The best-characterized scaffold of the ERK pathway is kinase suppressor of Ras 1 (KSR1). Initially identified as a suppressor of an activated Ras phenotype in Drosophila melanogaster and Caenorhabditis elegans,, KSR1 has a kinase domain with high homology with Raf-1 but mutations in residues critical for catalytic activity. Whether KSR1 has remaining kinase activity or whether it is a pseudokinase is still discussed in the literature., However, it is now accepted that the main function of KSR1 is as a scaffold of the ERK pathway, which regulates the intensity and duration of the ERK signal independent of catalytic function. KSR1 can interact with all kinases of the ERK pathway. MEK is constitutively bound, while Raf (Raf-1 or B-Raf) and ERK are recruited to KSR1 upon mitogen stimulation., However, KSR1 only binds less than 5% of endogenous Raf-1, indicating that KSR1 affects only a subset of Raf functions, and Raf members might be present in other protein complexes. KSR1 can activate Drosophila melanogaster Raf (which is closely related to mammalian B-Raf) allosterically by dimerization. In addition, KSR1 facilitates N-region phosphorylation of Raf-1 and B-Raf by recruiting CK2. Thus, KSR does not only regulate substrate availability but also catalytic activity, suggesting complex kinetic effects. In the context of cancer biology, KSR1 regulates Ras-mediated signaling, in particular differentiation, proliferation, and cellular transformation.- Gene deletion of KSR1 in the mouse had little effect on viability but decreased the oncogenic effects of the polyoma virus middle T antigen and blunted oncogenic Ras-mediated tumorigenesis., KSR1 depletion also accelerated the immortalization of cells and led to resistance to cisplatin-meditated apoptosis, while overexpression of KSR1 sensitized tumor cells to this anticancer agent and other drugs., Taken together, the functions of KSR1 in the context of Ras/ERK signaling vary dramatically and depend, like other scaffold proteins, on the level of expression. At low and physiological levels, KSR1 seems to work as a positive regulator of signaling. By contrast, overexpression of KSR1 has an inhibitory function on the activation of the ERK cascade., Recently, KSR2, a homolog of KSR1, was shown to participate in the calcium-mediated activation of ERK. This regulation is exerted by the calcium-dependent phosphatase calcineurin, which binds to KSR2 and dephosphorylates it, resulting in increased membrane translocation and ERK signaling. Another proteomic study showed that KSR2 preferentially interacts with A-Raf in response to TNF-α. The biological relevance of this interaction remains to be elucidated, but as KSR2 knockout mice have a striking metabolic dysregulation that includes obesity,, it is tempting to speculate that KSR2 may be involved in mediating A-Raf effects on metabolism.
Another group of Raf scaffolds is the connector enhancer of KSR (CNK) family of proteins. First identified as a modifier of KSR signaling in Drosophila melanogaster dCNK, mammals possess 3 isoforms that lack kinase activity but feature different protein-protein interaction domains that can bind a variety of client proteins including Raf-1 and B-Raf., Thus, CNK proteins seem to be superscaffolds that may integrate different signaling pathways. It is beyond the scope of this review to discuss CNK function in detail, and hence, we only will focus on the role of CNKs in Raf regulation. In Drosophila melanogaster, a multiprotein complex formed between dCNK, Raf, KSR, and a small adaptor protein HYP mediates Ras-induced activation of Raf., HYP has no mammalian homolog, and mammalian CNKs lack the Raf regulatory domain found in dCNK, suggesting a different mode of regulation. Mammalian CNK2 participates in the NGF-induced sustained ERK activation that is required for the neuronal differentiation of PC12 cells. The mechanism was not defined but, by analogy to CNK1, may involve facilitation of Raf activation. CNK1 can augment Raf-1 activation by increasing tyrosine phosphorylation of the N-region through recruiting c-Src. Interestingly, CNK1 also can bind RASSF1A and enhance apoptosis in a MST1/2-dependent manner. Thus, it is tempting to speculate that CNK1 may play a role in balancing apoptosis and proliferation by coordinating MST2 binding to RASSF1A and Raf-1, respectively.
In addition to KSR1, other ERK pathway scaffolds were implicated in tumor progression. Among them are the IQ motif containing GTPase-activating proteins (IQGAPs), a family of multidomain proteins., IQGAP1 directly interacts and modulates the functions of B-Raf, MEK, and ERK., Furthermore, IQGAP1 is required for the activation of B-Raf by EGF. As a result, IQGAP1 increases proliferation and reduces cellular differentiation. Thus, it comes as no surprise that IQGAP1 is involved in carcinogenesis., Augmented expression of IQGAPs was reported for several malignancies including cancers of the stomach, colon, lung, and prostate. Overexpression of IQGAP1 in human breast epithelial cells increased the formation and invasion of tumors, whereas reducing IQGAP1 expression had the opposite effect. Therefore, IGGAP1 is considered as a putative oncogene. Furthermore, IQGAPs are linked with metastasis, as IQGAP1 promotes cell migration and invasion via direct interactions with Cdc42, Rac1, actin, and calmodulin.
Prohibitin (PHB) facilitates the displacement of 14-3-3 from Raf-1 by activated Ras, thereby promoting plasma membrane localization and phosphorylation of Raf-1 at the activating S338. Interestingly, PHB binds only to Raf-1, but not to Ras, and may function as a chaperone of Raf to enable interaction with Ras. In the context of cancer, PHB function was connected to immortalization, aging, and cell cycle regulation. Furthermore, PHB overexpression was reported in breast cancer, human endometrial adenocarcinoma, gastric cancer, and bladder cancer, although it is not known if this is related to aberrant Raf function.
Scaffolds as Spatial Regulators of the ERK Pathway
Scaffold proteins are also crucial for the localization of the members of the ERK pathway to different subcellular signaling platforms. One such scaffold is similar expression to FGF (Sef-1), also known as interleukin-17 receptor (IL-17RD), which is situated at the Golgi apparatus. This transmembrane protein binds to activated MEK and facilitates activation of ERK but prevents ERK translocation to the nucleus. Therefore, ERK can only activate cytosolic targets. Interestingly, loss of Sef-1 expression is associated with high-grade metastatic prostate cancer. In clathrin-coated pits, the β-arrestins were proposed to augment ERK activation by scaffolding Raf-1, MEK, and ERK., The β-arrestins seem to act in a similar fashion as Sef-1, preventing ERK nuclear translocation and therefore restricting Ras signaling to the cytoplasmic effectors of the pathway.
The small scaffold MEK partner-1 (MP1) is an obligatory heterodimer with p14, and this complex interacts with MEK and ERK, targeting them to late endosomes., Recent results suggest that an additional adaptor, p18, is involved in specifying this subcellular localization.In vitro results indicated that MP1 also may facilitate MEK activation by B-Raf, although the mechanism is unknown. While decreased MP1 levels reduce ERK activation, overexpression of MP1 increases the binding of ERK to MEK and thus enhances the efficiency of ERK signaling., The speculation that the specific localization directed by MP1 generates signaling specificity was confirmed by the finding that MP1 specifically regulates PAK1-mediated ERK activation during cell adhesion and spreading but is not required for ERK activation by PDGF. Enhanced MP1 expression in several melanoma cell lines could be linked with a genetic translocation (4q23), suggesting a mechanism for the enhanced MAPK signaling in melanomas. In addition, MP1 may target MEK-ERK to high molecular weight protein complexes. Such complexes may be organized by MAPK organizer 1 (MORG1), a member of the WD-40 protein family, which was identified as an interaction partner of MP1 as well as Raf-1, B-Raf, MEK, and ERK. There is evidence that MP1 and MORG1 are part of a larger network built from nested scaffolds, as MEK binding to MORG1 is stabilized by MP1, Raf-1, and ERK. MORG1 acts like a classic scaffold with enhanced activation of ERK at low concentrations and being inhibitory at higher concentrations. Furthermore, MORG1 promotes ERK activity in response to serum or other signals. Interestingly, MORG1 was also shown to act as a scaffold with hypoxia-inducible factor prolyl hydroxylase 3 (PHD3) and downregulation of MORG1-augmented HIF-1 activity, suggesting MORG1 as a connection to other signaling networks.
The multidomain protein paxillin is a component of focal adhesions, providing a structural and signaling link between the actin cytoskeleton and the extracellular matrix (ECM). Paxillin constitutively interacts with MEK but in response to growth factors also binds to activated Raf and ERK, directing activated ERK to sites at the focal adhesions. The most significant impact of paxillin is on developmental processes and on tissue morphogenesis,- but it also plays a role in tumor cell invasion. Elevated levels of paxillin, together with enhanced Src activity, contribute to the high metastatic potential of human osteosarcomas, and in gastric cancer, high levels of paxillin correlated with advanced tumor stage and invasiveness.
Modulators of Protein Interactions in the ERK Pathway
Raf kinase inhibitor protein (RKIP) is receiving sharply increasing attention as a modulator of the ERK pathway and several other signaling pathways including G protein signaling and NFκB signaling.,- Initially identified as phosphatidylethanolamine-binding protein-1 (PEBP-1), RKIP was later identified as a negative modulator of Raf-1. RKIP binds Raf-1, MEK, and ERK. While RKIP can bind MEK and ERK simultaneously, binding to Raf-1 and MEK is mutually exclusive, disrupting the Raf-1–MEK complex and activation of MEK by Raf-1. RKIP also interferes with Raf-1 activation by preventing the interaction with PAK1 and Src kinases and the phosphorylation of the N-region. In this study, B-Raf activation was not affected. However, another study found that RKIP inhibited B-Raf activation in cells as well as its ability to phosphorylate MEK in vitro. Interestingly, in an RKIP-related protein, hPEBP4, the Raf- and MEK-binding sites, which overlap in RKIP, are separated by an insertion converting hPEBP4 into a scaffold for the Raf-1–MEK complex. Consequently, hPEBP4 enhances the activity of the ERK pathway in growing human myoblasts. Upon induction of differentiation, hPEBP4 expression rises to levels that exceed the optimal stoichiometric relationship to its client proteins and contribute to the inhibition of the ERK pathway observed during myoblast differentiation. This regulation highlights not only that scaffolds can assume both stimulating and inhibitory roles under physiological conditions but also that a primary function of such proteins is the fine tuning of the activation kinetics of signaling pathways. RKIP has a main function in causing switchlike activation behavior of ERK and in supporting oscillations of the pathway caused by the negative feedback from ERK to Ras activation. Physiologically, RKIP plays a major role as a suppressor of cancer invasiveness and metastasis in various cancers including common cancers of the prostate, breast, colon, and liver.
Other negative modulators of Raf-mediated signaling are members of the Sprouty (Spry) family, comprising various Spry and Spred (Spry-related proteins with an EVH1 domain) isoforms.- These proteins are negative feedback regulators of the ERK signaling pathway, and their expression is regulated by the cascade. Depending on the context, Spry proteins inhibit ERK signaling by binding and sequestering the Grb2-SOS complex, thus preventing Ras activation. Additionally, both Sprouty and SPRED can physically interact with Raf-1 and B-Raf, interfering with the phosphorylation of Raf on activating sites.- Spry genes and proteins were shown to be deregulated in different tumor types. Spry1 and Spry2 are downregulated in breast, prostate, and liver cancer., This downregulation seems to be due to hypermethylation of the promoter region, indicating that Sprys are putative tumor suppressors.
Genetic Alterations in Raf Family Genes
Raf Mutations in Cancer
For almost 2 decades, research focused on Raf-1 as the critical Ras effector of the Raf family. However, this changed when Davies et al. described mutations of B-Raf in 66% of melanomas and at a lower frequency in a wide range of human solid cancers. Further research revealed that approximately 2% of human malignancies carry a mutation in B-Raf, with highest frequencies observed in melanoma and carcinomas of the colon, thyroid gland, ovary, and biliary tract., Currently, more than 100 different B-Raf mutations were described, with V600E (formerly labeled as V599E) being by far the predominant lesion (Catalogue of Somatic Mutations in Cancer: www.sanger.ac.uk/genetics/CGP/cosmic). Most of the B-Raf mutants associated with cancer are located in exons 11 or 15 in the kinase domain. The biggest group of mutations (including V600E) affects residues that normally stabilize the kinase in the inactive form. Mutations of these amino acids disrupt this conformation, usually resulting in a significantly increased B-Raf kinase activity that leads to the constitutive activation of the ERK pathway. However, even impaired kinase activity mutants can constitutively activate the ERK pathway because of their ability to heterodimerize with and activate Raf-1., Interestingly, B-Raf mutations normally do not coexist with oncogenic mutations in Ras in human tumors, arguing that they are equivalent in their transforming effects. This conclusion indeed highlights B-Raf as a critical effector of Ras in cell transformation and cancer. In further support of this interpretation, mutant B-Raf has also been shown to be a critical step in tumorigenesis in mouse models of melanoma., Melanocyte-specific, conditional expression of the V600E mutation inserted into the endogenous BRAF gene locus resulted in the development of both benign nevi and malignant melanoma. This is in line with observations from human tumors, where B-Raf V600E is detected in approximately 44% of melanoma cases, but with even higher frequency in benign nevi, which do not progress into a malignant melanoma. This dormancy is probably due to the potent induction of senescence by B-Raf V600E, which suppresses tumorigenesis., These data suggest that additional secondary alterations cooperating with mutant B-Raf may be required. Further support of this hypothesis comes from observations demonstrating that B-Raf V600E induces senescence and needs secondary events like p16INK4A loss to overcome it. Interestingly, melanomas in the above-mentioned mouse model did not show any alterations in p16INK4A, suggesting that the pathogenetic mechanisms differ between the mouse and human or else that there are other, hitherto unknown possibilities for secondary genetic events. However, there are also B-Raf mutations associated with human cancer, which display impaired kinase activity, with D594V being the most frequent one. These mutations require Raf-1 to activate the ERK pathway, relying on the ability of catalytically compromised B-Raf to activate Raf-1 by heterodimerization., As Raf heterodimerization is augmented by activated Ras, these low-activity B-Raf mutants (in contrast to the high-activity mutants) can be found coexpressed with mutant Ras.
Although quite rare, cancer-associated mutations also were reported in Raf-1. They were first described in a mouse model of chemically induced lung cancer, in human cancer cell lines, and finally in patients with therapy-related acute myeloid leukemia (t-AML). Interestingly, the latter mutations were detected in the germline of affected patients but still exhibited weakly transforming and antiapoptotic properties. Therefore, they might constitute a hereditary predisposition to solid neoplasms and t-AML. This hypothesis was supported by the observation that constitutive activation of the ERK pathway in affected patients was only observed in malignant but not in the surrounding normal tissues. Indeed, further studies identified a leukemia-specific, somatic loss of RKIP as a genetic second hit that further promoted malignant transformation in these patients.
Besides mutations, other alterations of Raf genes were described in human malignancies as well. Rearrangements and fusions of B-Raf and Raf-1 to a variety of other genes have been described in thyroid cancer, pilocytic astrocytoma, prostate cancer, gastric cancer, and melanoma.- They seem to be particularly frequent in sporadic pilocytic astrocytoma, with more than 60% of cases demonstrating B-Raf rearrangements. Usually, the resulting fusion products lose the regulatory N-terminal region but retain an intact Raf kinase domain. They can activate the ERK pathway, transform transfected cell lines, and induce tumors in nude mice, underlining their functional role in the pathogenesis of human malignancies. Mutated B-Raf further was found amplified in human melanoma with a gain of chromosome 7q. This amplification is a frequent event in melanoma, suggesting that B-Raf mutations are one of the factors driving its selection. Elevated levels of A-Raf mRNA and protein were observed in a number of malignancies including head and neck squamous cell carcinomas and colon carcinomas. This study also showed that the splice factor heterogeneous nuclear ribonucleoprotein H (hnRNP H) is required for the correct splicing and expression of full-length A-Raf. Elevated expression of A-Raf was found in testicular germ cell tumor–derived cell lines caused by the duplication of the X chromosome.
Raf Mutations in Developmental Syndromes
Raf mutations are not only critical steps for tumorigenesis but also for the pathogenesis of rare developmental disorders, such as neurofibromatosis type 1 and Costello, Noonan, LEOPARD, and cardiofaciocutaneous (CFC) syndromes, which are reviewed in detail elsewhere.- Affected individuals present with overlapping yet distinct phenotypes that include a variable degree of mental retardation, cardiac defects, facial dysmorphisms, short stature, macrocephaly, and skin abnormalities. Germline mutations in Raf were first linked to these disorders when 2 groups simultaneously reported mutations in B-Raf causing CFC., The B-Raf mutations in CFC comprised mainly hitherto unknown mutations, which were more widely distributed across B-Raf as compared to their counterparts detected in human cancer. However, a few mutations are shared between cancer and CFC. Some of the CFC germline mutations resulted in increased B-Raf kinase activity and constitutive activation of the ERK pathway. Compared to the V600E mutation, the kinase activity and transforming capacity of CFC B-Raf mutants seem to be lower.,
Germline mutations in Raf-1 were recently described in both Noonan and LEOPARD syndromes., As with B-Raf, one of the mutations (S427G) was found in both Noonan syndrome and human malignancies, and again, some of the germline variants increase Raf-1 kinase activity and transforming ability., One might expect that carrying an oncogene in the germline increases the risk for the development of malignancies. Indeed, patients with neurofibromatosis type 1, Costello syndrome, and Noonan syndrome are at increased risk for developing a wide range of solid tumors and hematological malignancies.- Whether CFC and LEOPARD syndromes result in a predisposition to tumor development is an open question. The numbers of patients affected by these disorders are too low for a thorough statistical analysis, and descriptions of patients developing a malignant disorder are limited to case reports.- However, close monitoring of patients with all germline Ras/MAPK disorders, including LEOPARD and CFC syndromes, for the occurrence of neoplasias is often suggested.
Targeting Raf and MEK for Cancer Treatment
The efforts to develop drugs targeting the Raf family and their downstream effectors were increased after strategies implemented to inhibit Ras signaling failed in the preclinical and clinical studies. Different approaches included the inhibition of Raf kinase activity by small molecule inhibitors, decreasing Raf protein levels using antisense oligonucleotides, and targeting Raf protein-protein interactions, especially the Raf-Ras interaction. The first drugs were developed against Raf-1, but the discovery of B-Raf–activating mutation in tumors shifted the efforts toward the inhibition of this protein and of MEK1/2.
Raf Inhibitors
Sorafenib (BAY 43-9006) was the first Raf inhibitor to progress into clinical trials. Preclinical studies showed that sorafenib inhibited Raf-1 and B-Raf in tumor cell lines and xenograft models for Ras-dependent tumors., Today, sorafenib is approved for the treatment of advanced renal cell carcinoma (RCC) and unresectable hepatocellular carcinoma (HCC). However, sorafenib monotherapy failed to be clinically effective against other tumors such as melanoma, although it increased progression-free survival in combination with other treatments., Sorafenib was developed as a specific Raf-1 kinase inhibitor, and while it poorly inhibits mutant B-Raf, it is highly effective against several other kinases such as VEGF and PDGF receptors. In fact, sorafenib is now regarded as a multikinase inhibitor, and the success in HCC and RCC is probably primarily because of the inhibition of VEGF and PDGF receptors rather than Raf kinases. The results of the sorafenib clinical trials raised questions about the suitability of Raf proteins as therapeutic targets and the low predictive properties of the preclinical models. One possible explanation for the unexpected therapeutic target spectrum of sorafenib is that it is a poor B-Raf inhibitor. This conclusion led to the development of a new generation of inhibitors active against B-Raf. RAF265 is active against all Raf isoforms, mutant B-Raf, and VEGF-2. RAF265 inhibits cell proliferation in mutant B-Raf and N-Ras melanoma cells but has no effect in cell lines that express the normal genes. This finding led to the initiation of a series of clinical trials for the treatment of advanced melanoma in which patients were evaluated for B-Raf and N-Ras mutation before treatment. The results from these clinical trials are eagerly awaited and are expected to be published shortly. XL281 is another pan-Raf inhibitor that is active against mutant B-Raf and currently is in phase I clinical trials. Finally, PLX4032 is a potent B-Raf V600E selective kinase inhibitor that suppresses the activation of the ERK pathway and cell proliferation in melanoma xenografts., PLX4032 does not inhibit the ERK pathway in cells that do not express mutant B-Raf. A recent clinical phase I study reported a spectacular response rate in 81% of melanoma patients with mutant B-Raf. These results demonstrate that mutant B-Raf is an excellent therapeutic target in melanoma. However, this success did not come without cost, as 31% of patients developed skin tumors, keratoacanthomas, and squamous cell carcinomas. This side effect may be due to the enhancement of aberrant ERK pathway activation by drug-induced B-Raf–Raf-1 dimerization as discussed below. Although the skin tumors can be easily recognized and surgically removed, the appearance of malignancies in other less accessible organs remains a concern, and further studies are advocated before PLX4032 is approved for the treatment of metastatic melanomas.
The generation of drugs that target Raf interactions with other proteins is not as advanced but conceptually promising. MCP-110 is a small molecule that inhibits the Raf-Ras interaction. This compound decreased anchorage-independent growth in cell lines expressing oncogenic Ras but did not affect cell lines with a constitutively active Raf-1, indicating that MCP-110 is working specifically at the level of the Ras-Raf interaction. However, this agent did not progress into preclinical development. Another Raf interaction that was targeted is the binding to the Rb tumor suppressor protein. As mentioned above, Raf-1 was reported to bind to and phosphorylate Rb, resulting in Rb inhibition and S phase progression. Small synthetic peptides that interrupt the Raf-1–Rb interaction in cell lines suppressed the growth of A549 xenograft tumors. The use of peptides as drugs is limited by their short half-life and problems in their delivery, but similar results were obtained using the small-molecule drug RRD-251. This drug inhibited cell proliferation in vitro and suppressed the growth of xenograft tumors as well as tumor angiogenesis in a manner dependent on the expression of intact Rb. Although no clinical data are available, these results indicate that targeting the Raf-1–Rb interaction may be a successful antitumoral strategy. It also suggests that targeting other Raf interactions may be a good strategy. Of special interest would be the disruption of Raf-1 binding to Rok-α, which may promote the differentiation of epidermal skin tumor cells, and the dissociation of Raf-1 from MST1/2 and ASK1, which should activate the proapoptotic potential of these kinases. More interestingly, the disruption of the Raf-1/A-Raf–MST1/2 complex should compensate for the frequent loss of the RASSF1A tumor suppressor, which normally promotes the disruption of the MST–Raf-1 complex and activation of MST1/2. Such drugs would restore a natural tumor suppressor function and hence may be expected to be specific, efficacious, and without severe side effects.
A greater advance was made in targeting RAF expression using antisense oligonucleotides. The use of oligonucleotides for therapy is restricted by their susceptibility to degradation by nucleases, but the substitution of oxygen for sulfur in the phosphodiester linkages confers stability to these molecules. Using these chemical modifications, different compounds were generated, such as ISIS 5132 and ISIS 2503. ISIS 5132 is a 20-base phosphorothiate antisense oligonucleotide against Raf-1 that inhibited tumor progression in clinical trials. However, subsequent phase II clinical trials demonstrated that this compound was of no benefit as a single agent, and hence, it was not further developed. Another Raf-1–directed antisense agent that entered clinical trials is LErafAON, a liposome-entrapped derivative of a 15-mer antisense oligonucleotide. Packaging the antisense oligonucleotide within liposomes was expected to protect the oligonucleotide against nucleases and increase cell delivery. Unfortunately, several phase I clinical trials demonstrated a lack of objective responses but adverse side effects due to the liposomal formulation., The failure of the antisense drugs suspended the development of this approach for Raf inhibition. However, improvements in delivery methods, and the observation that small interfering RNA (siRNA) against mutant B-Raf, and Raf-1 can inhibit cell proliferation in melanoma and breast cancer xenograft models, respectively, may rekindle the interest in this strategy.
Paradoxical Effects of Raf Inhibitors
A decade ago, a report was published showing that the Raf inhibitor ZM 336372 produced a massive paradoxical activation of Raf kinases when cells were treated with the inhibitor, and then, Raf kinases were isolated and their activity measured in vitro. Now, 3 publications finally shed light on this apparent contradiction., The key is Raf-1 homodimerization or heterodimerization with B-Raf, which is driven by mutant Ras and facilitated by the Raf inhibitor drugs. The activation conferred by dimerization is not compromised if the kinase activity of one of the Raf dimerization partners is destroyed and the kinase activity of Raf-1–B-Raf heterodimers is higher than that of Raf-1 homodimers. Dimerization, induced by Ras or by Raf inhibitors, of mutant B-Raf V600E and Raf-1 actually dampens the overall kinase activity. These constellations exacerbate the activation effect when B-Raf–specific inhibitors are used and Raf dimerization is ushered by Ras mutations. Thus, Raf inhibitors are rather effective when B-Raf is mutated but ineffective when Ras is mutated., Beyond this shared theme, the 3 studies differ in mechanistic details.
In cell lines expressing high-activity B-Raf mutants, Raf inhibitors function as expected and efficiently reduced signaling through the ERK pathway. Consequently, cell proliferation was inhibited in vitro as well as in xenografts when cells were treated with B-Raf inhibitors. Surprisingly, in cell lines lacking an activating B-Raf mutation and expressing mutant Ras, the effects were opposite., Specific B-Raf inhibitors, such as PLX4720, enhanced ERK phosphorylation, and the cells demonstrated a high drug tolerance in xenograft models and proliferation assays. Moreover, the inhibitors initiated Raf heterodimerization, which enhanced the kinase activity and downstream signaling. Pan-Raf and less specific inhibitors, on the other hand, enhanced ERK signaling at a lower concentration, while higher doses reverted this effect and caused inhibition. These data suggest that both Raf-1 and B-Raf kinase activities present in a dimer need to be inhibited in order to shut down the signaling efficiently. Consistently, the inhibitor-induced ERK activity is Raf-1 dependent when B-Raf–specific inhibitors are used., Interestingly, B-Raf inhibitors are able to activate Raf-1 independently of B-Raf. PLX4720 robustly induces ERK phosphorylation in B-Raf–/– fibroblasts, which led to the conclusion that association with the inhibitor and drug-induced Raf-1 homodimerization is sufficient to enhance Raf kinase activity and ERK signaling. Heidorn et al. could also demonstrate in a mouse model that mutant K-Ras and kinase-dead B-Raf cooperated in the induction of melanomas. This is a worrying discovery, and it would be interesting to know if B-Raf inhibitors would have the same effect as kinase-dead B-Raf in promoting melanoma in a mutant K-Ras model.
In light of these data, the development of keratoacanthomas and squamous cell carcinoma in 31% of melanoma patients treated with PLX4720 may be ascribed to the paradoxical activation of Raf signaling by drug-induced heterodimerization. These results have direct implications for clinical practice. Firstly, the patient population, which should receive B-Raf–specific inhibitors, has to be carefully selected, as only tumors expressing activating B-Raf mutations will respond to the treatment, while mutant Ras expression may even worsen the condition. Furthermore, a combination therapy including pan-Raf and MEK inhibitors should also be considered in order to enhance the efficacy (see below) and limit adverse side effects and the fast onset of drug resistance.
MEK Inhibitors
The first highly selective and effective inhibitors against the ERK pathway were directed against MEK1/2. Work from different groups in preclinical models indicated that MEK inhibitors are highly effective if the pathway is activated by B-Raf mutations but rather ineffective against cells harboring mutant Ras.,CI-1040, the first MEK inhibitor to proceed to clinical phase I trials, had limited side effects and reduced ERK phosphorylation in more than 65% of the tumor biopsies. However, a subsequent phase II trial did not show clinical responses in a variety of tumor entities, and consequently, the development was stopped. Nevertheless, these studies suggested that targeting MEK was relatively safe, and new generations of MEK inhibitors were developed.PD0325901, one of these new-generation MEK inhibitors, is 50 times more potent than CI-1040 and has better pharmacological properties. However, the results from 2 clinical trials were disappointing, showing a high level of toxicity, and PD0325901 was dropped from further development. Currently, 7 MEK inhibitors are in different phases of clinical trials, and many more are in preclinical development. The results published so far show different antitumoral efficacies and also different levels of toxicity.
Thus, the development of drugs against the Raf family and MEK has produced both successes and disappointments, which have helped to develop new strategies that may lead to better compounds or better use of compounds. Most of the Raf and MEK inhibitors show limited effect as single agents, but they may still be effective in combination with classic chemotherapeutic agents or drugs that specifically target other signaling pathways, such as PI3K or growth factor receptor inhibitors. Deriving effective drug combinations will be helped by the rational prediction of drug responses using mathematical and computational modeling (see below) and by the identification of better biomarkers that allow discriminating, which patients would benefit from the treatment with Raf or MEK inhibitors, leading to the creation of personalized protocols for individual patients.
A Systems Biology View of the Ras-Raf Signaling Network
Despite the 25 years of research on Ras and Ras effectors, we are still bewildered by the diverse functionalities of this pathway, which can specify a multitude of often contradictory biological outcomes. While we have identified a large number of components and thoroughly characterized the individual functions of many of them, we still lack an understanding of how the Ras-Raf network processes signals to generate specific biological responses. Emerging evidence shows that much of this specificity is encoded by the pathway structure and dynamic changes in the connections between the proteins., These so-called emergent properties are difficult to understand by experimentation alone. Therefore, mathematical and computational modeling approaches were developed that allow us to analyze and predict network responses including the behavior of the Ras-Raf pathway.,
The Cascade Structure of Ras-Raf Signaling Networks: Amplification of Signals and Sensitivity
The “backbone” of the Raf pathway consists of a 3-tiered kinase cascade. This design allows for a larger repertoire of regulation by feedback, crosstalk, and scaffolding to increase the number of signaling functions a single pathway might perform. It also allows for signal amplification, turning a low-abundance, noisy signal into a higher abundance, clearer signal at each tier of the cascade. Measured in terms of active molecules, the amplification factor from active Ras to active ERK can be 6- to 30-fold.- Not only does this design amplify signals, but it also amplifies the sensitivity of the cascade output (ppERK) to the cascade input (RasGTP)., This property was also shown experimentally in Xenopus oocytes, where the effective cooperativity (Hill coefficient) increases with each tier of the cascade, so that a defined increase in the stimulation of the first component causes successively larger increases downstream. The activation mechanism by multisite phosphorylation adds further to this sensitivity amplification, if the phosphorylations occur in a distributive manner, where each phosphorylation requires a separate binding event between kinase and substrate.,In vitro ERK phosphorylation by MEK and dephosphorylation by MAPK phosphatase 3 (MPK3) follow such a distributive mechanism,, which based on theoretical considerations can give rise to switchlike behavior, bistability, and oscillations., While bistability of ERK activity was experimentally observed in several settings,, oscillations may be confined to certain experimental conditions and pathway topologies.,- Bistability means that the system is either “off” or “on,” even after the initiating stimulus has ceased. Thus, bistability is advantageous in biological processes that require irreversible decisions, such as cell cycle progression, differentiation, or cell death. In contrast, the biological role of oscillations of activities in signaling pathways is less obvious but may facilitate the temporal synchronization of processes.
Feedback Mechanisms: Tuning the Dynamics and Input/Output Sensitivity
The properties arising from signal and sensitivity amplification provide advantages, for instance, the ability to respond to small input signals, while filtering out noise through the switchlike behavior. However, there are also tradeoffs. Amplifiers may overshoot the desired output levels, and the more sensitive the cascade becomes, the quicker the output will saturate, and it will respond to a smaller range of inputs. Biology has found ways to avoid these tradeoffs, and they include feedback mechanisms. Modeling is very useful in analyzing how feedback loops change the behavior of a biological system, as it is usually difficult to isolate these feedbacks in an experimental setting but easy to do so within a mathematical model. The Ras/ERK pathway features a number of negative and positive feedback loops (Fig. 6), which can generate a wide variety of dynamic behavior.
Schematic diagram of feedback mechanisms in the Ras-ERK pathway. Short- and long-term negative feedbacks are colored light and dark blue, respectively. Short- and long-term positive feedbacks are colored orange and red, respectively.
Multiple layers and roles of negative feedback
The light blue lines in Figure 6 denote short-term negative feedback loops, which begin to act nearly immediately upon activation of ERK. Activated ERK leads to the phosphorylation and inactivation of the RasGEF SOS,, and modeling suggests that several phosphorylation sites on SOS all independently mediate strong negative feedback. ERK feedback phosphorylation affects targets at each upstream activation step. The first is the EGFR. Phosphorylation of T669 by ERK triggers multiple effects including decreases in receptor internalization, kinase activity, and the phosphorylation of selected substrates., Blocking this feedback inhibition by MEK inhibitors increased EGFR activity and enhanced epithelial-to-mesenchymal transition and migration. The next layer of negative feedback targets comprises adaptor protein working immediately downstream of the EGFR, that is, the Grb2-SOS complex,, and Gab1, a scaffolding protein involved in PI3K and RasGEF recruitment to the plasma membrane. Further downstream, Raf-1,, B-Raf,, and MEK are phosphorylated, leading to decreased pathway activity as discussed above.
When strong negative feedback is combined with the amplifier, constituted by the Raf-MEK-ERK cascade, the pathway adopts characteristics of a negative feedback amplifier (NFA). This circuitry is widely employed in electronic systems to confer response linearization and robustness to noise. These properties also exist in the biological NFA, as predicted by mathematical modeling and experimental validation. First, the NFA rendered ERK activation to become more linear in response to input dose, countering the effects of multiple kinase tiers and multisite phosphorylations that amplify input/output sensitivity and cause switchlike behavior. Second, ERK activation became robust to internal perturbations; for example, when MEK activity was incompletely inhibited, the attenuation of the negative feedback permitted the input to rise and sustain MEK activity. Breaking the negative feedback, for example, by expression of Raf-1 mutants that are resistant to feedback phosphorylation by ERK, very effectively sensitized the system to MEK inhibitors. Several important conclusions emanated from the analysis of the NFA design. Proteins embedded in NFAs are difficult drug targets because the NFA design will buffer any inhibition that is not complete. In contrast, inhibition of inputs outside of the NFA module, such as growth factor receptors, produced a linear dose response curve. Even more interestingly, weakening the negative feedback by a Raf inhibitor drastically improved the efficacy of a MEK inhibitor. Thus, the mathematical model made a concrete suggestion for a highly efficient drug combination, that is, Raf and MEK inhibitors, which from the experimentalists’ point of view appears so counterintuitive that it probably never would have been tested. Thus, the analysis of design principles of signaling pathways by mathematical models can give very applicable results.
The next layer in time is the delayed negative feedbacks that emanate from ERK but that are mediated by the transcriptional induction of feedback inhibitors. They are depicted by the dark blue lines in Figure 6 and only begin to take effect approximately 30 minutes or longer after ERK activation. Active ERK induces the transcription of multiple cytoplasmic and nuclear dual-specificity phosphatase (DUSP) isoforms, which dephosphorylate and deactivate ERK., In some instances, the stability and/or the phosphatase activity of DUSPs is also controlled by ERK activity, resulting in a positive feedforward loop embedded into this negative feedback., In addition, ERK stimulates the transcription of other pathway inhibitors, such as the Sprouty and Spred proteins,, which inhibit EGFR endocytosis, RasGEF recruitment, and Raf activation, and Mig6/RALT, which not only inhibits the activity of various receptor tyrosine kinases (RTKs), but also leads to increased ErbB1 receptor degradation in a manner apparently independent from the traditional ligand-stimulated pathway.
Another function of negative feedback is adaptation or return of ERK activity at steady state to near prestimulus levels despite the persistence of stimulus., Which negative feedback(s) are responsible for adaptation was the topic of many theoretical studies, but there is still no consensus as to which are the most important in general or if other mechanisms such as receptor downregulation play a role. When ERK itself is responsible for the direct negative feedback, then the system may adapt but will not exhibit perfect adaptation, where Ras and ERK activity returns exactly to prestimulus levels. The perfect adaptive behavior is characteristic of an engineering design termed “integral negative feedback,” where the strength of the negative feedback is proportional to the time-integrated ERK activity. Recent modeling work suggests that the long-term, transcriptional negative feedbacks might act as such integral negative feedback circuits, as mRNA responses of active ERK-dependent genes are proportional to the total time active ERK spends in the nucleus. Thus, a function of the delayed negative feedback distinct from the short-term negative feedbacks may be to achieve adaptation, that is, a resetting of the system after the response has been executed. Adaptation of biochemical networks usually requires either a negative feedback that acts proportional to the input or a negative feedback combined with a positive feedforward (incoherent feedforward loop).,
Positive feedback flips the switches
The orange and red lines in Figure 6 denote short-term and long-term positive feedbacks, respectively. The short-term feedbacks include the ERK-mediated 1) phosphorylation and inactivation of RKIP, a protein that inhibits Raf’s ability to activate MEK; 2) phosphorylation of PEA-15, which releases ERK from this cytosolic anchor protein and allows the nuclear accumulation of ppERK,; and 3) phosphorylation of NF-1, a RasGAP whose ability to convert RasGTP back into the inactive RasGDP form is inhibited by ERK phosphorylation. In addition, RasGTP produced by the RasGEF SOS can bind to another site in SOS that allosterically stimulates GEF activity. In T lymphocytes, this positive feedback is key to establish a bistable Ras response that contributes to the establishment of memory cells, which maintain the ability to be rapidly reactivated by the specific antigens they have encountered before. The biological relevance of positive feedback includes the generation of bistability. Bistability typically is brought about by strong positive feedback, which maintains the response even after the input was removed. Thus, bistability in the Ras/ERK pathway underlies processes that require clear and sustained “on” and “off” signals, such as memory formation in individual neurons and cell fate decisions of neuronal, and lymphoid cells., The long-term positive feedbacks involve the ERK-induced autocrine production or release of growth factors that entertain further ERK activity by stimulating receptors. Such autocrine mechanisms are widely implicated in development, cell differentiation, and tumorigenesis.-
A Brief History of Crosstalk: Integrating Various Signals
Although we know that the Ras/ERK pathway is only part of a much larger network, most of our analysis treats it as an isolated entity. This approach has proven very effective, but we have to be aware that much of the distinction between pathways may simply reflect the historic sequence of discoveries that conveniently helps us to compartmentalize the network into conceptually and experimentally accessible entities. From this vantage point, we usually summarize connections between pathways as crosstalk. Much of the known crosstalk between ERK and other pathways is positive, but there are also modes of negative crosstalk. The assortment of crosstalks discussed here is certainly incomplete, and many of the crosstalk mechanisms are likely to be cell type dependent. However, a striking observation is that many crosstalk mechanisms are under the control of Ras or act on Ras, suggesting that Ras plays a major role in coordinating this crosstalk.
A prime example is PI3K, which can be activated by receptors and by Ras directly. PI3K phosphorylates PIP(4,5)2 to produce PIP(3,4,5)3, which binds with high affinity to proteins containing pleckstrin homology (PH) domains, such as the Gab scaffolds. This allows PI3K to facilitate activation of Ras through 2 mechanisms, as Gab1 recruits SOS complexes and also the phosphatase SHP2. SOS directly activates Ras, while SHP2 maintains RasGTP levels by dephosphorylating residues on RTKs that recruit RasGAP. Furthermore, PI3K also leads to activation of PAK, which phosphorylates Raf-1 on the activating S338 residue. A main effector of PI3K is Akt, which intersects with the ERK pathway on several levels. Akt was reported to inhibit Raf-1 by phosphorylation of S259, which however was not substantiated in subsequent work., More interestingly, Akt shares several substrates with ERK, where phosphorylation by ERK and PI3K acts synergistically. Examples include the proapoptotic protein BAD, which is jointly inactivated by Akt phosphorylation and ERK-dependent phosphorylation. Similarly, both Akt and ERK jointly phosphorylate PEA-15, a cytosolic anchor for ERK, thereby inducing the release of ERK and allowing ERK phosphorylation and nuclear accumulation. In this case, positive crosstalk is coupled with positive feedback, which is likely to yield highly nonlinear synergistic effects. Another example for positive crosstalk emanates from the PLCγ pathway, which is activated by many RTKs on the same time scale as Ras. Active PLCγ cleaves the phospholipid PIP(4,5)2 to produce diacylglycerol (DAG) that stays in the plasma membrane and soluble inositol triphosphate (IP3) that induces a calcium release from the endoplasmic reticulum (ER). DAG and calcium, alone and in combination, can activate 2 classes of proteins important for Ras/ERK signaling: a family of RasGEFs called the RasGRPs and various protein kinase C (PKC) isoforms. There is not yet evidence of whether the RasGRPs are subjected to negative feedback regulation as is SOS. However, RasGRP primes SOS for allosteric activation by RasGTP and thereby plays an essential role to cause the bistable Ras activation in T lymphocytes discussed above. However, calcium via its binding protein calmodulin also can inhibit Ras-mediated activation of ERK signaling., Activated PKC enhances signaling through the ERK pathway through several mechanisms including the inhibition of RKIP by phosphorylation on S153., RKIP can also be disabled by ERK phosphorylation on S99, and this event plays a major role in the crosstalk between the ERK and Wnt pathways.
What is the purpose of all this crosstalk? Likely, the primary reason is the integration of various environmental cues by the cell to make appropriate decisions. For instance, it is known that under normal circumstances, adherent cells will not proliferate while not attached, and the crosstalk between adhesion/PI3K signaling and ERK signaling may coordinate this response., Similarly, the crosstalk between the cAMP and ERK systems, which was discussed above, can inhibit Raf-1 but stimulate B-Raf. Thus, the specific response depends on the expression of Raf-1 and B-Raf, which can differ between cell types and tissues.
Conclusions
What have we learned about the Raf-MEK-ERK pathway as an effector of Ras in the 17 years since this relationship was brought to light? A simple summary could be that Rafs are the “first in, last out” in Ras signaling. In accounting, this is a method of inventory valuation, which is based on the assumption that the cost of goods purchased first is the cost of goods sold last. In other words, in a growing enterprise, the value of the old stock rises with the value of new additions. Undoubtedly, exploring the Ras signaling network is a blooming industry. We now have entered a phase in which the mapping of the components of signaling networks is rapidly progressing and for the Ras network may soon approach completion. However, the result is a telephone book full of names rather than an ordnance survey type of map that in detail connects the names with pathway topologies. We also need to be able to trace the wanderers’ steps in time to understand the function of the pathways. Here again, the old stock is taking the lead in developing approaches to draw the missing lines, track the temporal relationships, and analyze the composite behavior of simple modules embedded in complex networks. The most important insights will be those that transcend the directory of names and provide an understanding of the design principles of biological systems and their evolution. This will enable us to address the next big challenge: How do biochemical signaling networks generate biological specificity? The current state of analysis of the Ras-Raf network offers a glimpse into this new world. The concepts and tools developed in this process hopefully will widen this glimpse into a window overlooking the whole Ras network.
Acknowledgments
The authors apologize to those whose works could not be cited because of space constraints.
Footnotes
The author(s) declared no potential conflicts of interest with respect to the authorship and/or publication of this article.
This work was supported by the Science Foundation Ireland [grant number 06/CE/B1129]; an EMBO long-term fellowship (A.Z.); a Marie Curie Fellowship [number 236758 (M.B.)]; and the European Union Seventh Framework Programme (FP7) [grant number LSHC-CT-2006-037731 (D.M.)].
References
1. Rapp UR, Goldsborough MD, Mark GE, et al. Structure and biological activity of v-raf, a unique oncogene transduced by a retrovirus. Proc Natl Acad Sci U S A. 1983;80:4218-22 [PMC free article] [PubMed] [Google Scholar]
2. Sutrave P, Bonner TI, Rapp UR, Jansen HW, Patschinsky T, Bister K. Nucleotide sequence of avian retroviral oncogene v-mil: homologue of murine retroviral oncogene v-raf. Nature. 1984;309:85-8 [PubMed] [Google Scholar]
3. Moelling K, Heimann B, Beimling P, Rapp UR, Sander T. Serine- and threonine-specific protein kinase activities of purified gag-mil and gag-raf proteins. Nature. 1984;312:558-61 [PubMed] [Google Scholar]
4. Bonner TI, Kerby SB, Sutrave P, Gunnell MA, Mark G, Rapp UR. Structure and biological activity of human homologs of the raf/mil oncogene. Mol Cell Biol. 1985;5:1400-7 [PMC free article] [PubMed] [Google Scholar]
5. Jansen HW, Bister K. Nucleotide sequence analysis of the chicken gene c-mil, the progenitor of the retroviral oncogene v-mil. Virology. 1985;143:359-67 [PubMed] [Google Scholar]
6. Wasylyk C, Wasylyk B, Heidecker G, Huleihel M, Rapp UR. Expression of raf oncogenes activates the PEA1 transcription factor motif. Mol Cell Biol. 1989;9:2247-50 [PMC free article] [PubMed] [Google Scholar]
7. Kolch W, Heidecker G, Lloyd P, Rapp UR. Raf-1 protein kinase is required for growth of induced NIH/3T3 cells. Nature. 1991;349:426-8 [PubMed] [Google Scholar]
8. Jamal S, Ziff E. Transactivation of c-fos and beta-actin genes by raf as a step in early response to transmembrane signals. Nature. 1990;344:463-6 [PubMed] [Google Scholar]
9. Kyriakis JM, App H, Zhang XF, et al. Raf-1 activates MAP kinase-kinase. Nature. 1992;358:417-21 [PubMed] [Google Scholar]
10. Dent P, Haser W, Haystead TA, Vincent LA, Roberts TM, Sturgill TW. Activation of mitogen-activated protein kinase kinase by v-Raf in NIH 3T3 cells and in vitro. Science. 1992;257:1404-7 [PubMed] [Google Scholar]
11. Zhang XF, Settleman J, Kyriakis JM, et al. Normal and oncogenic p21ras proteins bind to the amino-terminal regulatory domain of c-Raf-1. Nature. 1993;364:308-13 [PubMed] [Google Scholar]
12. Warne PH, Viciana PR, Downward J. Direct interaction of Ras and the amino-terminal region of Raf-1 in vitro. Nature. 1993;364:352-5 [PubMed] [Google Scholar]
13. Vojtek AB, Hollenberg SM, Cooper JA. Mammalian Ras interacts directly with the serine/threonine kinase Raf. Cell. 1993;74:205-14 [PubMed] [Google Scholar]
14. Van Aelst L, Barr M, Marcus S, Polverino A, Wigler M. Complex formation between RAS and RAF and other protein kinases. Proc Natl Acad Sci U S A. 1993;90:6213-7 [PMC free article] [PubMed] [Google Scholar]
15. Moodie SA, Willumsen BM, Weber MJ, Wolfman A. Complexes of Ras.GTP with Raf-1 and mitogen-activated protein kinase kinase. Science. 1993;260:1658-61 [PubMed] [Google Scholar]
16. Chen J, Fujii K, Zhang L, Roberts T, Fu H. Raf-1 promotes cell survival by antagonizing apoptosis signal-regulating kinase 1 through a MEK-ERK independent mechanism. Proc Natl Acad Sci U S A. 2001;98:7783-8 [PMC free article] [PubMed] [Google Scholar]
17. O’Neill E, Rushworth L, Baccarini M, Kolch W. Role of the kinase MST2 in suppression of apoptosis by the proto-oncogene product Raf-1. Science. 2004;306:2267-70 [PubMed] [Google Scholar]
18. Piazzolla D, Meissl K, Kucerova L, Rubiolo C, Baccarini M. Raf-1 sets the threshold of Fas sensitivity by modulating Rok-alpha signaling. J Cell Biol. 2005;171:1013-22 [PMC free article] [PubMed] [Google Scholar]
19. Ehrenreiter K, Piazzolla D, Velamoor V, et al. Raf-1 regulates Rho signaling and cell migration. J Cell Biol. 2005;168:955-64 [PMC free article] [PubMed] [Google Scholar]
20. Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949-54 [PubMed] [Google Scholar]
21. Wellbrock C, Karasarides M, Marais R. The RAF proteins take centre stage. Nat Rev Mol Cell Biol. 2004;5:875-85 [PubMed] [Google Scholar]
22. Leicht DT, Balan V, Kaplun A, et al. Raf kinases: function, regulation and role in human cancer. Biochim Biophys Acta. 2007;1773:1196-212 [PMC free article] [PubMed] [Google Scholar]
23. Dhillon AS, Hagan S, Rath O, Kolch W. MAP kinase signalling pathways in cancer. Oncogene. 2007;26:3279-90 [PubMed] [Google Scholar]
24. Kholodenko BN, Hancock JF, Kolch W. Signalling ballet in space and time. Nat Rev Mol Cell Biol. 2010;11:414-26 [PMC free article] [PubMed] [Google Scholar]
25. Tran NH, Wu X, Frost JA. B-Raf and Raf-1 are regulated by distinct autoregulatory mechanisms. J Biol Chem. 2005;280:16244-53 [PubMed] [Google Scholar]
26. Dhillon AS, Meikle S, Yazici Z, Eulitz M, Kolch W. Regulation of Raf-1 activation and signalling by dephosphorylation. EMBO J. 2002;21:64-71 [PMC free article] [PubMed] [Google Scholar]
27. Chong H, Lee J, Guan KL. Positive and negative regulation of Raf kinase activity and function by phosphorylation. EMBO J. 2001;20:3716-27 [PMC free article] [PubMed] [Google Scholar]
28. Nassar N, Horn G, Herrmann C, Scherer A, McCormick F, Wittinghofer A. The 2.2 A crystal structure of the Ras-binding domain of the serine/threonine kinase c-Raf1 in complex with Rap1A and a GTP analogue. Nature. 1995;375:554-60 [PubMed] [Google Scholar]
29. Emerson SD, Madison VS, Palermo RE, et al. Solution structure of the Ras-binding domain of c-Raf-1 and identification of its Ras interaction surface. Biochemistry. 1995;34:6911-8 [PubMed] [Google Scholar]
30. Mott HR, Carpenter JW, Zhong S, Ghosh S, Bell RM, Campbell SL. The solution structure of the Raf-1 cysteine-rich domain: a novel ras and phospholipid binding site. Proc Natl Acad Sci U S A. 1996;93:8312-7 [PMC free article] [PubMed] [Google Scholar]

31. Wan PT, Garnett MJ, Roe SM, et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell. 2004;116:855-67 [PubMed] [Google Scholar]
32. Hatzivassiliou G, Song K, Yen I, et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature. 2010;464:431-5 [PubMed] [Google Scholar]
33. Cutler RE, Jr., Stephens RM, Saracino MR, Morrison DK. Autoregulation of the Raf-1 serine/threonine kinase. Proc Natl Acad Sci U S A. 1998;95:9214-9 [PMC free article] [PubMed] [Google Scholar]
34. Chong H, Guan KL. Regulation of Raf through phosphorylation and N terminus-C terminus interaction. J Biol Chem. 2003;278:36269-76 [PubMed] [Google Scholar]
35. Heidecker G, Huleihel M, Cleveland JL, et al. Mutational activation of c-raf-1 and definition of the minimal transforming sequence. Mol Cell Biol. 1990;10:2503-12 [PMC free article] [PubMed] [Google Scholar]
36. Yip-Schneider MT, Miao W, Lin A, Barnard DS, Tzivion G, Marshall MS. Regulation of the Raf-1 kinase domain by phosphorylation and 14-3-3 association. Biochem J. 2000;351:151-9 [PMC free article] [PubMed] [Google Scholar]
37. Tran NH, Frost JA. Phosphorylation of Raf-1 by p21-activated kinase 1 and Src regulates Raf-1 autoinhibition. J Biol Chem. 2003;278:11221-6 [PubMed] [Google Scholar]
38. Terai K, Matsuda M. Ras binding opens c-Raf to expose the docking site for mitogen-activated protein kinase kinase. EMBO Rep. 2005;6:251-5 [PMC free article] [PubMed] [Google Scholar]
39. Raabe T, Rapp UR. Ras signaling: PP2A puts Ksr and Raf in the right place. Curr Biol. 2003;13:R635-7 [PubMed] [Google Scholar]
40. Ory S, Zhou M, Conrads TP, Veenstra TD, Morrison DK. Protein phosphatase 2A positively regulates Ras signaling by dephosphorylating KSR1 and Raf-1 on critical 14-3-3 binding sites. Curr Biol. 2003;13:1356-64 [PubMed] [Google Scholar]
41. Kubicek M, Pacher M, Abraham D, Podar K, Eulitz M, Baccarini M. Dephosphorylation of Ser-259 regulates Raf-1 membrane association. J Biol Chem. 2002;277:7913-9 [PubMed] [Google Scholar]
42. Goetz CA, O’Neil JJ, Farrar MA. Membrane localization, oligomerization, and phosphorylation are required for optimal raf activation. J Biol Chem. 2003;278:51184-9 [PubMed] [Google Scholar]
43. Jaumot M, Hancock JF. Protein phosphatases 1 and 2A promote Raf-1 activation by regulating 14-3-3 interactions. Oncogene. 2001;20:3949-58 [PubMed] [Google Scholar]
44. Rommel C, Radziwill G, Lovric J, et al. Activated Ras displaces 14-3-3 protein from the amino terminus of c-Raf-1. Oncogene. 1996;12:609-19 [PubMed] [Google Scholar]
45. Kiel C, Serrano L. Cell type-specific importance of ras-c-raf complex association rate constants for MAPK signaling. Sci Signal. 2009;2:ra38. [PubMed] [Google Scholar]
46. Buday L, Warne PH, Downward J. Downregulation of the Ras activation pathway by MAP kinase phosphorylation of Sos. Oncogene. 1995;11:1327-31 [PubMed] [Google Scholar]
47. Dong C, Waters SB, Holt KH, Pessin JE. SOS phosphorylation and disassociation of the Grb2-SOS complex by the ERK and JNK signaling pathways. J Biol Chem. 1996;271:6328-32 [PubMed] [Google Scholar]
48. Matsunaga-Udagawa R, Fujita Y, Yoshiki S, et al. The scaffold protein Shoc2/SUR-8 accelerates the interaction of Ras and Raf. J Biol Chem. 2010;285:7818-26 [PMC free article] [PubMed] [Google Scholar]
49. Rodriguez-Viciana P, Oses-Prieto J, Burlingame A, Fried M, McCormick F. A phosphatase holoenzyme comprised of Shoc2/Sur8 and the catalytic subunit of PP1 functions as an M-Ras effector to modulate Raf activity. Mol Cell. 2006;22:217-30 [PubMed] [Google Scholar]
50. Williams JG, Drugan JK, Yi GS, Clark GJ, Der CJ, Campbell SL. Elucidation of binding determinants and functional consequences of Ras/Raf-cysteine-rich domain interactions. J Biol Chem. 2000;275:22172-9 [PubMed] [Google Scholar]
51. Bondeva T, Balla A, Varnai P, Balla T. Structural determinants of Ras-Raf interaction analyzed in live cells. Mol Biol Cell. 2002;13:2323-33 [PMC free article] [PubMed] [Google Scholar]
52. Roy S, Lane A, Yan J, McPherson R, Hancock JF. Activity of plasma membrane-recruited Raf-1 is regulated by Ras via the Raf zinc finger. J Biol Chem. 1997;272:20139-45 [PubMed] [Google Scholar]
53. Fischer A, Hekman M, Kuhlmann J, Rubio I, Wiese S, Rapp UR. B- and C-RAF display essential differences in their binding to Ras: the isotype-specific N terminus of B-RAF facilitates Ras binding. J Biol Chem. 2007;282:26503-16 [PubMed] [Google Scholar]
54. Kraft CA, Garrido JL, Fluharty E, Leiva-Vega L, Romero G. Role of phosphatidic acid in the coupling of the ERK cascade. J Biol Chem. 2008;283:36636-45 [PMC free article] [PubMed] [Google Scholar]
55. Hekman M, Hamm H, Villar AV, et al. Associations of B- and C-Raf with cholesterol, phosphatidylserine, and lipid second messengers: preferential binding of Raf to artificial lipid rafts. J Biol Chem. 2002;277:24090-102 [PubMed] [Google Scholar]
56. Andresen BT, Rizzo MA, Shome K, Romero G. The role of phosphatidic acid in the regulation of the Ras/MEK/Erk signaling cascade. FEBS Lett. 2002;531:65-8 [PubMed] [Google Scholar]
57. Johnson LM, James KM, Chamberlain MD, Anderson DH. Identification of key residues in the A-Raf kinase important for phosphoinositide lipid binding specificity. Biochemistry. 2005;44:3432-40 [PubMed] [Google Scholar]
58. Zhao C, Du G, Skowronek K, Frohman MA, Bar-Sagi D. Phospholipase D2-generated phosphatidic acid couples EGFR stimulation to Ras activation by Sos. Nat Cell Biol. 2007;9:706-12 [PubMed] [Google Scholar]
59. Ariotti N, Liang H, Xu Y, et al. Epidermal growth factor receptor activation remodels the plasma membrane lipid environment to induce nanocluster formation. Mol Cell Biol. 2010;30:3795-804 [PMC free article] [PubMed] [Google Scholar]
60. Inder K, Harding A, Plowman SJ, Philips MR, Parton RG, Hancock JF. Activation of the MAPK module from different spatial locations generates distinct system outputs. Mol Biol Cell. 2008;19:4776-84 [PMC free article] [PubMed] [Google Scholar]
61. Prior IA, Harding A, Yan J, Sluimer J, Parton RG, Hancock JF. GTP-dependent segregation of H-ras from lipid rafts is required for biological activity. Nat Cell Biol. 2001;3:368-75 [PubMed] [Google Scholar]
62. Tian T, Harding A, Inder K, Plowman S, Parton RG, Hancock JF. Plasma membrane nanoswitches generate high-fidelity Ras signal transduction. Nat Cell Biol. 2007;9:905-14 [PubMed] [Google Scholar]
63. Chiu VK, Bivona T, Hach A, et al. Ras signalling on the endoplasmic reticulum and the Golgi. Nat Cell Biol. 2002;4:343-50 [PubMed] [Google Scholar]
64. Matallanas D, Sanz-Moreno V, Arozarena I, et al. Distinct utilization of effectors and biological outcomes resulting from site-specific Ras activation: Ras functions in lipid rafts and Golgi complex are dispensable for proliferation and transformation. Mol Cell Biol. 2006;26:100-16 [PMC free article] [PubMed] [Google Scholar]
65. Daniels MA, Teixeiro E, Gill J, et al. Thymic selection threshold defined by compartmentalization of Ras/MAPK signalling. Nature. 2006;444:724-9 [PubMed] [Google Scholar]
66. Diaz B, Barnard D, Filson A, MacDonald S, King A, Marshall M. Phosphorylation of Raf-1 serine 338-serine 339 is an essential regulatory event for Ras-dependent activation and biological signaling. Mol Cell Biol. 1997;17:4509-16 [PMC free article] [PubMed] [Google Scholar]
67. Xiang X, Zang M, Waelde CA, Wen R, Luo Z. Phosphorylation of 338SSYY341 regulates specific interaction between Raf-1 and MEK1. J Biol Chem. 2002;277:44996-5003 [PubMed] [Google Scholar]
68. Edin ML, Juliano RL. Raf-1 serine 338 phosphorylation plays a key role in adhesion-dependent activation of extracellular signal-regulated kinase by epidermal growth factor. Mol Cell Biol. 2005;25:4466-75 [PMC free article] [PubMed] [Google Scholar]
69. Fabian JR, Daar IO, Morrison DK. Critical tyrosine residues regulate the enzymatic and biological activity of Raf-1 kinase. Mol Cell Biol. 1993;13:7170-9 [PMC free article] [PubMed] [Google Scholar]
70. Marais R, Light Y, Paterson HF, Marshall CJ. Ras recruits Raf-1 to the plasma membrane for activation by tyrosine phosphorylation. EMBO J. 1995;14:3136-45 [PMC free article] [PubMed] [Google Scholar]
71. Xia K, Mukhopadhyay NK, Inhorn RC, et al. The cytokine-activated tyrosine kinase JAK2 activates Raf-1 in a p21ras-dependent manner. Proc Natl Acad Sci U S A. 1996;93:11681-6 [PMC free article] [PubMed] [Google Scholar]
72. King AJ, Sun H, Diaz B, et al. The protein kinase Pak3 positively regulates Raf-1 activity through phosphorylation of serine 338. Nature. 1998;396:180-3 [PubMed] [Google Scholar]
73. Sun H, King AJ, Diaz HB, Marshall MS. Regulation of the protein kinase Raf-1 by oncogenic Ras through phosphatidylinositol 3-kinase, Cdc42/Rac and Pak. Curr Biol. 2000;10:281-4 [PubMed] [Google Scholar]
74. Chaudhary A, King WG, Mattaliano MD, et al. Phosphatidylinositol 3-kinase regulates Raf1 through Pak phosphorylation of serine 338. Curr Biol. 2000;10:551-4 [PubMed] [Google Scholar]
75. Chiloeches A, Mason CS, Marais R. S338 phosphorylation of Raf-1 is independent of phosphatidylinositol 3-kinase and Pak3. Mol Cell Biol. 2001;21:2423-34 [PMC free article] [PubMed] [Google Scholar]
76. Zang M, Gong J, Luo L, et al. Characterization of Ser338 phosphorylation for Raf-1 activation. J Biol Chem. 2008;283:31429-37 [PMC free article] [PubMed] [Google Scholar]
77. Ritt DA, Zhou M, Conrads TP, Veenstra TD, Copeland TD, Morrison DK. CK2 is a component of the KSR1 scaffold complex that contributes to Raf kinase activation. Curr Biol. 2007;17:179-84 [PubMed] [Google Scholar]
Ras Activation Of Raffle
78. Lorentzen A, Kinkhabwala A, Rocks O, Vartak N, Bastiaens PI. Regulation of Ras localization by acylation enables a mode of intracellular signal propagation. Sci Signal. 2010;3:ra68. [PubMed] [Google Scholar]
79. von Kriegsheim A, Pitt A, Grindlay GJ, Kolch W, Dhillon AS. Regulation of the Raf-MEK-ERK pathway by protein phosphatase 5. Nat Cell Biol. 2006;8:1011-6 [PubMed] [Google Scholar]
80. Rajakulendran T, Sahmi M, Lefrancois M, Sicheri F, Therrien M. A dimerization-dependent mechanism drives RAF catalytic activation. Nature. 2009;461:542-5 [PubMed] [Google Scholar]
81. Garnett MJ, Rana S, Paterson H, Barford D, Marais R. Wild-type and mutant B-RAF activate C-RAF through distinct mechanisms involving heterodimerization. Mol Cell. 2005;20:963-9 [PubMed] [Google Scholar]
82. Rushworth LK, Hindley AD, O’Neill E, Kolch W. Regulation and role of Raf-1/B-Raf heterodimerization. Mol Cell Biol. 2006;26:2262-72 [PMC free article] [PubMed] [Google Scholar]
83. Fan F, Feng L, He J, et al. RKTG sequesters B-Raf to the Golgi apparatus and inhibits the proliferation and tumorigenicity of human malignant melanoma cells. Carcinogenesis. 2008;29:1157-63 [PubMed] [Google Scholar]
84. Feng L, Xie X, Ding Q, et al. Spatial regulation of Raf kinase signaling by RKTG. Proc Natl Acad Sci U S A. 2007;104:14348-53 [PMC free article] [PubMed] [Google Scholar]
85. Zhu J, Balan V, Bronisz A, et al. Identification of Raf-1 S471 as a novel phosphorylation site critical for Raf-1 and B-Raf kinase activities and for MEK binding. Mol Biol Cell. 2005;16:4733-44 [PMC free article] [PubMed] [Google Scholar]
86. Zhang BH, Guan KL. Activation of B-Raf kinase requires phosphorylation of the conserved residues Thr598 and Ser601. EMBO J. 2000;19:5429-39 [PMC free article] [PubMed] [Google Scholar]
87. Harding A, Hsu V, Kornfeld K, Hancock JF. Identification of residues and domains of Raf important for function in vivo and in vitro. J Biol Chem. 2003;278:45519-27 [PubMed] [Google Scholar]
88. Park S, Rath O, Beach S, et al. Regulation of RKIP binding to the N-region of the Raf-1 kinase. FEBS Lett. 2006;580:6405-12 [PMC free article] [PubMed] [Google Scholar]
89. Rath O, Park S, Tang HH, et al. The RKIP (Raf-1 Kinase Inhibitor Protein) conserved pocket binds to the phosphorylated N-region of Raf-1 and inhibits the Raf-1-mediated activated phosphorylation of MEK. Cell Signal. 2008;20:935-41 [PubMed] [Google Scholar]
90. Yeung K, Janosch P, McFerran B, et al. Mechanism of suppression of the Raf/MEK/extracellular signal-regulated kinase pathway by the raf kinase inhibitor protein. Mol Cell Biol. 2000;20:3079-85 [PMC free article] [PubMed] [Google Scholar]
91. Yeung K, Seitz T, Li S, et al. Suppression of Raf-1 kinase activity and MAP kinase signalling by RKIP. Nature. 1999;401:173-7 [PubMed] [Google Scholar]
92. Dougherty MK, Muller J, Ritt DA, et al. Regulation of Raf-1 by direct feedback phosphorylation. Mol Cell. 2005;17:215-24 [PubMed] [Google Scholar]
93. Cirit M, Wang CC, Haugh JM. Systematic quantification of negative feedback mechanisms in the extracellular signal-regulated kinase (ERK) signaling network. J Biol Chem. 2010;285:36736-44 [PMC free article] [PubMed] [Google Scholar]
94. Balan V, Leicht DT, Zhu J, et al. Identification of novel in vivo Raf-1 phosphorylation sites mediating positive feedback Raf-1 regulation by extracellular signal-regulated kinase. Mol Biol Cell. 2006;17:1141-53 [PMC free article] [PubMed] [Google Scholar]
95. Baljuls A, Schmitz W, Mueller T, et al. Positive regulation of A-RAF by phosphorylation of isoform-specific hinge segment and identification of novel phosphorylation sites. J Biol Chem. 2008;283:27239-54 [PubMed] [Google Scholar]
96. Gerits N, Kostenko S, Shiryaev A, Johannessen M, Moens U. Relations between the mitogen-activated protein kinase and the cAMP-dependent protein kinase pathways: comradeship and hostility. Cell Signal. 2008;20:1592-607 [PubMed] [Google Scholar]
97. Wu J, Dent P, Jelinek T, Wolfman A, Weber MJ, Sturgill TW. Inhibition of the EGF-activated MAP kinase signaling pathway by adenosine 3′,5′-monophosphate. Science. 1993;262:1065-9 [PubMed] [Google Scholar]
98. Hafner S, Adler HS, Mischak H, et al. Mechanism of inhibition of Raf-1 by protein kinase A. Mol Cell Biol. 1994;14:6696-703 [PMC free article] [PubMed] [Google Scholar]
99. Dhillon AS, Pollock C, Steen H, Shaw PE, Mischak H, Kolch W. Cyclic AMP-dependent kinase regulates Raf-1 kinase mainly by phosphorylation of serine 259. Mol Cell Biol. 2002;22:3237-46 [PMC free article] [PubMed] [Google Scholar]
100. Dumaz N, Marais R. Protein kinase A blocks Raf-1 activity by stimulating 14-3-3 binding and blocking Raf-1 interaction with Ras. J Biol Chem. 2003;278:29819-23 [PubMed] [Google Scholar]
101. Mischak H, Seitz T, Janosch P, et al. Negative regulation of Raf-1 by phosphorylation of serine 621. Mol Cell Biol. 1996;16:5409-18 [PMC free article] [PubMed] [Google Scholar]
102. Dhillon AS, Yip YY, Grindlay GJ, et al. The C-terminus of Raf-1 acts as a 14-3-3-dependent activation switch. Cell Signal. 2009;21:1645-51 [PubMed] [Google Scholar]
103. Zimmermann S, Moelling K. Phosphorylation and regulation of Raf by Akt (protein kinase B). Science. 1999;286:1741-4 [PubMed] [Google Scholar]
104. Subramaniam S, Shahani N, Strelau J, et al. Insulin-like growth factor 1 inhibits extracellular signal-regulated kinase to promote neuronal survival via the phosphatidylinositol 3-kinase/protein kinase A/c-Raf pathway. J Neurosci. 2005;25:2838-52 [PubMed] [Google Scholar]
105. Garcia R, Grindlay J, Rath O, Fee F, Kolch W. Regulation of human myoblast differentiation by PEBP4. EMBO Rep. 2009;10:278-84 [PMC free article] [PubMed] [Google Scholar]
106. Marais R, Light Y, Paterson HF, Mason CS, Marshall CJ. Differential regulation of Raf-1, A-Raf, and B-Raf by oncogenic ras and tyrosine kinases. J Biol Chem. 1997;272:4378-83 [PubMed] [Google Scholar]
107. Weber CK, Slupsky JR, Herrmann C, Schuler M, Rapp UR, Block C. Mitogenic signaling of Ras is regulated by differential interaction with Raf isozymes. Oncogene. 2000;19:169-76 [PubMed] [Google Scholar]
108. Baljuls A, Mueller T, Drexler HC, Hekman M, Rapp UR. Unique N-region determines low basal activity and limited inducibility of A-RAF kinase: the role of N-region in the evolutionary divergence of RAF kinase function in vertebrates. J Biol Chem. 2007;282:26575-90 [PubMed] [Google Scholar]
109. Adams JA. Activation loop phosphorylation and catalysis in protein kinases: is there functional evidence for the autoinhibitor model?Biochemistry. 2003;42:601-7 [PubMed] [Google Scholar]
110. Yao Z, Seger R. The ERK signaling cascade: views from different subcellular compartments. Biofactors. 2009;35:407-16 [PubMed] [Google Scholar]
111. Yamamori B, Kuroda S, Shimizu K, Fukui K, Ohtsuka T, Takai Y. Purification of a Ras-dependent mitogen-activated protein kinase kinase kinase from bovine brain cytosol and its identification as a complex of B-Raf and 14-3-3 proteins. J Biol Chem. 1995;270:11723-6 [PubMed] [Google Scholar]
112. Hmitou I, Druillennec S, Valluet A, Peyssonnaux C, Eychene A. Differential regulation of B-raf isoforms by phosphorylation and autoinhibitory mechanisms. Mol Cell Biol. 2007;27:31-43 [PMC free article] [PubMed] [Google Scholar]
113. Brummer T, Martin P, Herzog S, Misawa Y, Daly RJ, Reth M. Functional analysis of the regulatory requirements of B-Raf and the B-Raf(V600E) oncoprotein. Oncogene. 2006;25:6262-76 [PubMed] [Google Scholar]
114. Konig S, Guibert B, Morice C, Vernier P, Barnier JV. Phosphorylation by PKA of a site unique to B-Raf kinase. C R Acad Sci III. 2001;324:673-81 [PubMed] [Google Scholar]
115. Qiu W, Zhuang S, von Lintig FC, Boss GR, Pilz RB. Cell type-specific regulation of B-Raf kinase by cAMP and 14-3-3 proteins. J Biol Chem. 2000;275:31921-9 [PubMed] [Google Scholar]
116. Okada T, Hu CD, Jin TG, Kariya K, Yamawaki-Kataoka Y, Kataoka T. The strength of interaction at the Raf cysteine-rich domain is a critical determinant of response of Raf to Ras family small GTPases. Mol Cell Biol. 1999;19:6057-64 [PMC free article] [PubMed] [Google Scholar]
117. Vossler MR, Yao H, York RD, Pan MG, Rim CS, Stork PJ. cAMP activates MAP kinase and Elk-1 through a B-Raf- and Rap1-dependent pathway. Cell. 1997;89:73-82 [PubMed] [Google Scholar]
118. Bos JL, de Bruyn K, Enserink J, et al. The role of Rap1 in integrin-mediated cell adhesion. Biochem Soc Trans. 2003;31:83-6 [PubMed] [Google Scholar]
119. Rodriguez-Viciana P, Sabatier C, McCormick F. Signaling specificity by Ras family GTPases is determined by the full spectrum of effectors they regulate. Mol Cell Biol. 2004;24:4943-54 [PMC free article] [PubMed] [Google Scholar]
120. Im E, von Lintig FC, Chen J, et al. Rheb is in a high activation state and inhibits B-Raf kinase in mammalian cells. Oncogene. 2002;21:6356-65 [PubMed] [Google Scholar]
121. Karbowniczek M, Robertson GP, Henske EP. Rheb inhibits C-raf activity and B-raf/C-raf heterodimerization. J Biol Chem. 2006;281:25447-56 [PubMed] [Google Scholar]
122. Yee WM, Worley PF. Rheb interacts with Raf-1 kinase and may function to integrate growth factor- and protein kinase A-dependent signals. Mol Cell Biol. 1997;17:921-33 [PMC free article] [PubMed] [Google Scholar]
123. Kitayama H, Sugimoto Y, Matsuzaki T, Ikawa Y, Noda M. A ras-related gene with transformation suppressor activity. Cell. 1989;56:77-84 [PubMed] [Google Scholar]
124. Cook SJ, Rubinfeld B, Albert I, McCormick F. RapV12 antagonizes Ras-dependent activation of ERK1 and ERK2 by LPA and EGF in Rat-1 fibroblasts. EMBO J. 1993;12:3475-85 [PMC free article] [PubMed] [Google Scholar]
125. Carey KD, Watson RT, Pessin JE, Stork PJ. The requirement of specific membrane domains for Raf-1 phosphorylation and activation. J Biol Chem. 2003;278:3185-96 [PubMed] [Google Scholar]
126. Xu TR, Vyshemirsky V, Gormand A, et al. Inferring signaling pathway topologies from multiple perturbation measurements of specific biochemical species. Sci Signal. 2010;3:ra20 [PubMed] [Google Scholar]
127. Awasthi A, Samarakoon A, Chu H, et al. Rap1b facilitates NK cell functions via IQGAP1-mediated signalosomes. J Exp Med. 2010;207:1923-38 [PMC free article] [PubMed] [Google Scholar]
128. Papin C, Barnier JV, Eychene A, Calothy G. [B-raf gene encodes for multiple isoforms with Mek-1 kinase activity]. C R Seances Soc Biol Fil. 1995;189:71-85 [PubMed] [Google Scholar]
129. Barnier JV, Papin C, Eychene A, Lecoq O, Calothy G. The mouse B-raf gene encodes multiple protein isoforms with tissue-specific expression. J Biol Chem. 1995;270:23381-9 [PubMed] [Google Scholar]
130. Yokoyama T, Takano K, Yoshida A, et al. DA-Raf1, a competent intrinsic dominant-negative antagonist of the Ras-ERK pathway, is required for myogenic differentiation. J Cell Biol. 2007;177:781-93 [PMC free article] [PubMed] [Google Scholar]
131. Nekhoroshkova E, Albert S, Becker M, Rapp UR. A-RAF kinase functions in ARF6 regulated endocytic membrane traffic. PLoS One. 2009;4:e4647. [PMC free article] [PubMed] [Google Scholar]
132. Dozier C, Ansieau S, Ferreira E, Coll J, Stehelin D. An alternatively spliced c-mil/raf mRNA is predominantly expressed in chicken muscular tissues and conserved among vertebrate species. Oncogene. 1991;6:1307-11 [PubMed] [Google Scholar]
133. Yaffe MB, Rittinger K, Volinia S, et al. The structural basis for 14-3-3:phosphopeptide binding specificity. Cell. 1997;91:961-71 [PubMed] [Google Scholar]
134. Fischer A, Baljuls A, Reinders J, et al. Regulation of RAF activity by 14-3-3 proteins: RAF kinases associate functionally with both homo- and heterodimeric forms of 14-3-3 proteins. J Biol Chem. 2009;284:3183-94 [PubMed] [Google Scholar]
135. Hekman M, Wiese S, Metz R, et al. Dynamic changes in C-Raf phosphorylation and 14-3-3 protein binding in response to growth factor stimulation: differential roles of 14-3-3 protein binding sites. J Biol Chem. 2004;279:14074-86 [PubMed] [Google Scholar]
136. Light Y, Paterson H, Marais R. 14-3-3 antagonizes Ras-mediated Raf-1 recruitment to the plasma membrane to maintain signaling fidelity. Mol Cell Biol. 2002;22:4984-96 [PMC free article] [PubMed] [Google Scholar]
137. Tzivion G, Luo Z, Avruch J. A dimeric 14-3-3 protein is an essential cofactor for Raf kinase activity. Nature. 1998;394:88-92 [PubMed] [Google Scholar]
138. Thorson JA, Yu LW, Hsu AL, et al. 14-3-3 proteins are required for maintenance of Raf-1 phosphorylation and kinase activity. Mol Cell Biol. 1998;18:5229-38 [PMC free article] [PubMed] [Google Scholar]
139. Moelling K, Schad K, Bosse M, Zimmermann S, Schweneker M. Regulation of Raf-Akt cross-talk. J Biol Chem. 2002;277:31099-106 [PubMed] [Google Scholar]
140. Guan KL, Figueroa C, Brtva TR, et al. Negative regulation of the serine/threonine kinase B-Raf by Akt. J Biol Chem. 2000;275:27354-9 [PubMed] [Google Scholar]
141. Cook SJ, McCormick F. Inhibition by cAMP of Ras-dependent activation of Raf. Science. 1993;262:1069-72 [PubMed] [Google Scholar]
142. Sprenkle AB, Davies SP, Carling D, Hardie DG, Sturgill TW. Identification of Raf-1 Ser621 kinase activity from NIH 3T3 cells as AMP-activated protein kinase. FEBS Lett. 1997;403:254-8 [PubMed] [Google Scholar]
143. Noble C, Mercer K, Hussain J, et al. CRAF autophosphorylation of serine 621 is required to prevent its proteasome-mediated degradation. Mol Cell. 2008;31:862-72 [PMC free article] [PubMed] [Google Scholar]
144. Ritt DA, Monson DM, Specht SI, Morrison DK. Impact of feedback phosphorylation and Raf heterodimerization on normal and mutant B-Raf signaling. Mol Cell Biol. 2010;30:806-19 [PMC free article] [PubMed] [Google Scholar]
145. MacNicol MC, Muslin AJ, MacNicol AM. Disruption of the 14-3-3 binding site within the B-Raf kinase domain uncouples catalytic activity from PC12 cell differentiation. J Biol Chem. 2000;275:3803-9 [PubMed] [Google Scholar]
146. Weber CK, Slupsky JR, Kalmes HA, Rapp UR. Active Ras induces heterodimerization of cRaf and BRaf. Cancer Res. 2001;61:3595-8 [PubMed] [Google Scholar]
147. Farrar MA, Alberol I, Perlmutter RM. Activation of the Raf-1 kinase cascade by coumermycin-induced dimerization. Nature. 1996;383:178-81 [PubMed] [Google Scholar]
148. Luo Z, Tzivion G, Belshaw PJ, Vavvas D, Marshall M, Avruch J. Oligomerization activates c-Raf-1 through a Ras-dependent mechanism. Nature. 1996;383:181-5 [PubMed] [Google Scholar]
149. Matheny SA, White MA. Signaling threshold regulation by the Ras effector IMP. J Biol Chem. 2009;284:11007-11 [PMC free article] [PubMed] [Google Scholar]
150. Chadee DN, Xu D, Hung G, et al. Mixed-lineage kinase 3 regulates B-Raf through maintenance of the B-Raf/Raf-1 complex and inhibition by the NF2 tumor suppressor protein. Proc Natl Acad Sci U S A. 2006;103:4463-8 [PMC free article] [PubMed] [Google Scholar]
151. Mason CS, Springer CJ, Cooper RG, Superti-Furga G, Marshall CJ, Marais R. Serine and tyrosine phosphorylations cooperate in Raf-1, but not B-Raf activation. EMBO J. 1999;18:2137-48 [PMC free article] [PubMed] [Google Scholar]
152. Hall-Jackson CA, Eyers PA, Cohen P, et al. Paradoxical activation of Raf by a novel Raf inhibitor. Chem Biol. 1999;6:559-68 [PubMed] [Google Scholar]
153. Poulikakos PI, Zhang C, Bollag G, Shokat KM, Rosen N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature. 2010;464:427-30 [PMC free article] [PubMed] [Google Scholar]
154. Heidorn SJ, Milagre C, Whittaker S, et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell. 2010;140:209-21 [PMC free article] [PubMed] [Google Scholar]
155. Papin C, Denouel A, Calothy G, Eychene A. Identification of signalling proteins interacting with B-Raf in the yeast two-hybrid system. Oncogene. 1996;12:2213-21 [PubMed] [Google Scholar]
156. Frost JA, Steen H, Shapiro P, et al. Cross-cascade activation of ERKs and ternary complex factors by Rho family proteins. EMBO J. 1997;16:6426-38 [PMC free article] [PubMed] [Google Scholar]
157. Eblen ST, Slack JK, Weber MJ, Catling AD. Rac-PAK signaling stimulates extracellular signal-regulated kinase (ERK) activation by regulating formation of MEK1-ERK complexes. Mol Cell Biol. 2002;22:6023-33 [PMC free article] [PubMed] [Google Scholar]
158. Eblen ST, Slack-Davis JK, Tarcsafalvi A, Parsons JT, Weber MJ, Catling AD. Mitogen-activated protein kinase feedback phosphorylation regulates MEK1 complex formation and activation during cellular adhesion. Mol Cell Biol. 2004;24:2308-17 [PMC free article] [PubMed] [Google Scholar]
159. Khosravi-Far R, Solski PA, Clark GJ, Kinch MS, Der CJ. Activation of Rac1, RhoA, and mitogen-activated protein kinases is required for Ras transformation. Mol Cell Biol. 1995;15:6443-53 [PMC free article] [PubMed] [Google Scholar]
160. Qiu RG, Chen J, Kirn D, McCormick F, Symons M. An essential role for Rac in Ras transformation. Nature. 1995;374:457-9 [PubMed] [Google Scholar]
161. Catalanotti F, Reyes G, Jesenberger V, et al. A Mek1-Mek2 heterodimer determines the strength and duration of the Erk signal. Nat Struct Mol Biol. 2009;16:294-303 [PubMed] [Google Scholar]
162. Hindley A, Kolch W. Extracellular signal regulated kinase (ERK)/mitogen activated protein kinase (MAPK)-independent functions of Raf kinases. J Cell Sci. 2002;115:1575-81 [PubMed] [Google Scholar]
163. Murakami MS, Morrison DK. Raf-1 without MEK?Sci STKE. 2001;2001:pe30. [PubMed] [Google Scholar]
164. Tan CM, Kelvin DJ, Litchfield DW, Ferguson SS, Feldman RD. Tyrosine kinase-mediated serine phosphorylation of adenylyl cyclase. Biochemistry. 2001;40:1702-9 [PubMed] [Google Scholar]
165. Ding Q, Gros R, Gray ID, Taussig R, Ferguson SS, Feldman RD. Raf kinase activation of adenylyl cyclases: isoform-selective regulation. Mol Pharmacol. 2004;66:921-8 [PubMed] [Google Scholar]
166. Beazely MA, Alan JK, Watts VJ. Protein kinase C and epidermal growth factor stimulation of Raf1 potentiates adenylyl cyclase type 6 activation in intact cells. Mol Pharmacol. 2005;67:250-9 [PubMed] [Google Scholar]
167. Wang S, Ghosh RN, Chellappan SP. Raf-1 physically interacts with Rb and regulates its function: a link between mitogenic signaling and cell cycle regulation. Mol Cell Biol. 1998;18:7487-98 [PMC free article] [PubMed] [Google Scholar]
168. Zheng L, Lee WH. The retinoblastoma gene: a prototypic and multifunctional tumor suppressor. Exp Cell Res. 2001;264:2-18 [PubMed] [Google Scholar]
169. Dasgupta P, Sun J, Wang S, et al. Disruption of the Rb--Raf-1 interaction inhibits tumor growth and angiogenesis. Mol Cell Biol. 2004;24:9527-41 [PMC free article] [PubMed] [Google Scholar]
170. Kinkade R, Dasgupta P, Carie A, et al. A small molecule disruptor of Rb/Raf-1 interaction inhibits cell proliferation, angiogenesis, and growth of human tumor xenografts in nude mice. Cancer Res. 2008;68:3810-8 [PMC free article] [PubMed] [Google Scholar]
171. Broustas CG, Grammatikakis N, Eto M, Dent P, Brautigan DL, Kasid U. Phosphorylation of the myosin-binding subunit of myosin phosphatase by Raf-1 and inhibition of phosphatase activity. J Biol Chem. 2002;277:3053-9 [PubMed] [Google Scholar]
172. Shimizu M, Wang W, Walch ET, Dunne PW, Epstein HF. Rac-1 and Raf-1 kinases, components of distinct signaling pathways, activate myotonic dystrophy protein kinase. FEBS Lett. 2000;475:273-7 [PubMed] [Google Scholar]
173. Pfleiderer P, Sumandea MP, Rybin VO, Wang C, Steinberg SF. Raf-1: a novel cardiac troponin T kinase. J Muscle Res Cell Motil. 2009;30:67-72 [PMC free article] [PubMed] [Google Scholar]
174. Galabova-Kovacs G, Kolbus A, Matzen D, et al. ERK and beyond: insights from B-Raf and Raf-1 conditional knockouts. Cell Cycle. 2006;5:1514-8 [PubMed] [Google Scholar]
175. Pritchard CA, Bolin L, Slattery R, Murray R, McMahon M. Post-natal lethality and neurological and gastrointestinal defects in mice with targeted disruption of the A-Raf protein kinase gene. Curr Biol. 1996;6:614-7 [PubMed] [Google Scholar]
176. Galabova-Kovacs G, Matzen D, Piazzolla D, et al. Essential role of B-Raf in ERK activation during extraembryonic development. Proc Natl Acad Sci U S A. 2006;103:1325-30 [PMC free article] [PubMed] [Google Scholar]
177. Mikula M, Schreiber M, Husak Z, et al. Embryonic lethality and fetal liver apoptosis in mice lacking the c-raf-1 gene. EMBO J. 2001;20:1952-62 [PMC free article] [PubMed] [Google Scholar]
178. Huser M, Luckett J, Chiloeches A, et al. MEK kinase activity is not necessary for Raf-1 function. EMBO J. 2001;20:1940-51 [PMC free article] [PubMed] [Google Scholar]
179. Rubiolo C, Piazzolla D, Meissl K, et al. A balance between Raf-1 and Fas expression sets the pace of erythroid differentiation. Blood. 2006;108:152-9 [PubMed] [Google Scholar]
180. Kolbus A, Pilat S, Husak Z, et al. Raf-1 antagonizes erythroid differentiation by restraining caspase activation. J Exp Med. 2002;196:1347-53 [PMC free article] [PubMed] [Google Scholar]
181. Chang F, Steelman LS, Shelton JG, et al. Regulation of cell cycle progression and apoptosis by the Ras/Raf/MEK/ERK pathway [review]. Int J Oncol. 2003;22:469-80 [PubMed] [Google Scholar]
182. Baumann B, Weber CK, Troppmair J, et al. Raf induces NF-kappaB by membrane shuttle kinase MEKK1, a signaling pathway critical for transformation. Proc Natl Acad Sci U S A. 2000;97:4615-20 [PMC free article] [PubMed] [Google Scholar]
183. Bertrand F, Philippe C, Antoine PJ, et al. Insulin activates nuclear factor kappa B in mammalian cells through a Raf-1-mediated pathway. J Biol Chem. 1995;270:24435-41 [PubMed] [Google Scholar]
184. Troppmair J, Hartkamp J, Rapp UR. Activation of NF-kappa B by oncogenic Raf in HEK 293 cells occurs through autocrine recruitment of the stress kinase cascade. Oncogene. 1998;17:685-90 [PubMed] [Google Scholar]
185. Norris JL, Baldwin AS., Jr Oncogenic Ras enhances NF-kappaB transcriptional activity through Raf-dependent and Raf-independent mitogen-activated protein kinase signaling pathways. J Biol Chem. 1999;274:13841-6 [PubMed] [Google Scholar]
186. Galmiche A, Fueller J, Santel A, et al. Isoform-specific interaction of C-RAF with mitochondria. J Biol Chem. 2008;283:14857-66 [PubMed] [Google Scholar]
187. Zhong J, Troppmair J, Rapp UR. Independent control of cell survival by Raf-1 and Bcl-2 at the mitochondria. Oncogene. 2001;20:4807-16 [PubMed] [Google Scholar]
188. Majewski M, Nieborowska-Skorska M, Salomoni P, et al. Activation of mitochondrial Raf-1 is involved in the antiapoptotic effects of Akt. Cancer Res. 1999;59:2815-9 [PubMed] [Google Scholar]
189. Wang HG, Rapp UR, Reed JC. Bcl-2 targets the protein kinase Raf-1 to mitochondria. Cell. 1996;87:629-38 [PubMed] [Google Scholar]
190. Wu X, Carr HS, Dan I, Ruvolo PP, Frost JA. p21 activated kinase 5 activates Raf-1 and targets it to mitochondria. J Cell Biochem. 2008;105:167-75 [PMC free article] [PubMed] [Google Scholar]
191. Jin S, Zhuo Y, Guo W, Field J. p21-activated kinase 1 (Pak1)-dependent phosphorylation of Raf-1 regulates its mitochondrial localization, phosphorylation of BAD, and Bcl-2 association. J Biol Chem. 2005;280:24698-705 [PubMed] [Google Scholar]
192. von Gise A, Lorenz P, Wellbrock C, et al. Apoptosis suppression by Raf-1 and MEK1 requires MEK- and phosphatidylinositol 3-kinase-dependent signals. Mol Cell Biol. 2001;21:2324-36 [PMC free article] [PubMed] [Google Scholar]
193. Fueller J, Becker M, Sienerth AR, Fischer A, Hotz C, Galmiche A. C-RAF activation promotes BAD poly-ubiquitylation and turn-over by the proteasome. Biochem Biophys Res Commun. 2008;370:552-6 [PubMed] [Google Scholar]
194. Hindley A, Kolch W. Raf-1 and B-Raf promote protein kinase C theta interaction with BAD. Cell Signal. 2007;19:547-55 [PubMed] [Google Scholar]
195. Le Mellay V, Troppmair J, Benz R, Rapp UR, et al. Negative regulation of mitochondrial VDAC channels by C-Raf kinase. BMC Cell Biol. 2002;3:14. [PMC free article] [PubMed] [Google Scholar]
196. Tobiume K, Matsuzawa A, Takahashi T, et al. ASK1 is required for sustained activations of JNK/p38 MAP kinases and apoptosis. EMBO Rep. 2001;2:222-8 [PMC free article] [PubMed] [Google Scholar]
197. Alavi AS, Acevedo L, Min W, Cheresh DA. Chemoresistance of endothelial cells induced by basic fibroblast growth factor depends on Raf-1-mediated inhibition of the proapoptotic kinase, ASK1. Cancer Res. 2007;67:2766-72 [PubMed] [Google Scholar]
198. Yamaguchi O, Watanabe T, Nishida K, et al. Cardiac-specific disruption of the c-raf-1 gene induces cardiac dysfunction and apoptosis. J Clin Invest. 2004;114:937-43 [PMC free article] [PubMed] [Google Scholar]
199. Praskova M, Khoklatchev A, Ortiz-Vega S, Avruch J. Regulation of the MST1 kinase by autophosphorylation, by the growth inhibitory proteins, RASSF1 and NORE1, and by Ras. Biochem J. 2004;381:453-62 [PMC free article] [PubMed] [Google Scholar]
200. Matallanas D, Romano D, Yee K, et al. RASSF1A elicits apoptosis through an MST2 pathway directing proapoptotic transcription by the p73 tumor suppressor protein. Mol Cell. 2007;27:962-75 [PMC free article] [PubMed] [Google Scholar]
201. Zhang X, Milton CC, Humbert PO, Harvey KF. Transcriptional output of the Salvador/warts/hippo pathway is controlled in distinct fashions in Drosophila melanogaster and mammalian cell lines. Cancer Res. 2009;69:6033-41 [PubMed] [Google Scholar]
202. Pan D. The hippo signaling pathway in development and cancer. Dev Cell. 2010;19:491-505 [PMC free article] [PubMed] [Google Scholar]
203. Zender L, Spector MS, Xue W, et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell. 2006;125:1253-67 [PMC free article] [PubMed] [Google Scholar]
204. Avruch J, Zhou D, Fitamant J, Bardeesy N. Mst1/2 signalling to Yap: gatekeeper for liver size and tumour development. Br J Cancer. 2011;104:24-32 [PMC free article] [PubMed] [Google Scholar]
205. Basu S, Totty NF, Irwin MS, Sudol M, Downward J. Akt phosphorylates the Yes-associated protein, YAP, to induce interaction with 14-3-3 and attenuation of p73-mediated apoptosis. Mol Cell. 2003;11:11-23 [PubMed] [Google Scholar]
206. Strano S, Monti O, Pediconi N, et al. The transcriptional coactivator Yes-associated protein drives p73 gene-target specificity in response to DNA Damage. Mol Cell. 2005;18:447-59 [PubMed] [Google Scholar]
207. Bertini E, Oka T, Sudol M, Strano S, Blandino G. YAP: at the crossroad between transformation and tumor suppression. Cell Cycle. 2009;8:49-57 [PubMed] [Google Scholar]
208. Avruch J, Xavier R, Bardeesy N, et al. Rassf family of tumor suppressor polypeptides. J Biol Chem. 2009;284:11001-5 [PMC free article] [PubMed] [Google Scholar]
209. Richter AM, Pfeifer GP, Dammann RH. The RASSF proteins in cancer: from epigenetic silencing to functional characterization. Biochim Biophys Acta. 2009;1796:114-28 [PubMed] [Google Scholar]
210. Romano D, Matallanas D, Weitsman G, Preisinger C, Ng T, Kolch W. Proapoptotic kinase MST2 coordinates signaling crosstalk between RASSF1A, Raf-1, and Akt. Cancer Res. 2010;70:1195-203 [PMC free article] [PubMed] [Google Scholar]
211. Nilsson JA, Cleveland JL. Myc pathways provoking cell suicide and cancer. Oncogene. 2003;22:9007-21 [PubMed] [Google Scholar]
212. O’Neill E, Kolch W. Taming the Hippo: Raf-1 controls apoptosis by suppressing MST2/Hippo. Cell Cycle. 2005;4:365-7 [PubMed] [Google Scholar]
213. Niault T, Sobczak I, Meissl K, et al. From autoinhibition to inhibition in trans: the Raf-1 regulatory domain inhibits Rok-alpha kinase activity. J Cell Biol. 2009;187:335-42 [PMC free article] [PubMed] [Google Scholar]
214. Ehrenreiter K, Kern F, Velamoor V, et al. Raf-1 addiction in Ras-induced skin carcinogenesis. Cancer Cell. 2009;16:149-60 [PubMed] [Google Scholar]
215. Rauch J, O’Neill E, Mack B, et al. Heterogeneous nuclear ribonucleoprotein H blocks MST2-mediated apoptosis in cancer cells by regulating A-Raf transcription. Cancer Res. 2010;70:1679-88 [PMC free article] [PubMed] [Google Scholar]
216. Rauch J, Ahlemann M, Schaffrik M, et al. Allogenic antibody-mediated identification of head and neck cancer antigens. Biochem Biophys Res Commun. 2004;323:156-62 [PubMed] [Google Scholar]
217. Le Mellay V, Houben R, Troppmair J, et al. Regulation of glycolysis by Raf protein serine/threonine kinases. Adv Enzyme Regul. 2002;42:317-32 [PubMed] [Google Scholar]
218. Christofk HR, Vander Heiden MG, Harris MH, et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature. 2008;452:230-3 [PubMed] [Google Scholar]
219. Mazurek S, Drexler HC, Troppmair J, Eigenbrodt E, Rapp UR. Regulation of pyruvate kinase type M2 by A-Raf: a possible glycolytic stop or go mechanism. Anticancer Res. 2007;27:3963-71 [PubMed] [Google Scholar]
220. Buchakjian MR, Kornbluth S. The engine driving the ship: metabolic steering of cell proliferation and death. Nat Rev Mol Cell Biol. 2010;11:715-27 [PubMed] [Google Scholar]
221. Dehmelt L, Bastiaens PI. Spatial organization of intracellular communication: insights from imaging. Nat Rev Mol Cell Biol. 2010;11:440-52 [PubMed] [Google Scholar]
222. Kolch W, Pitt A. Functional proteomics to dissect tyrosine kinase signalling pathways in cancer. Nat Rev Cancer. 2010;10:618-29 [PubMed] [Google Scholar]
223. McKay MM, Morrison DK. Integrating signals from RTKs to ERK/MAPK. Oncogene. 2007;26:3113-21 [PubMed] [Google Scholar]
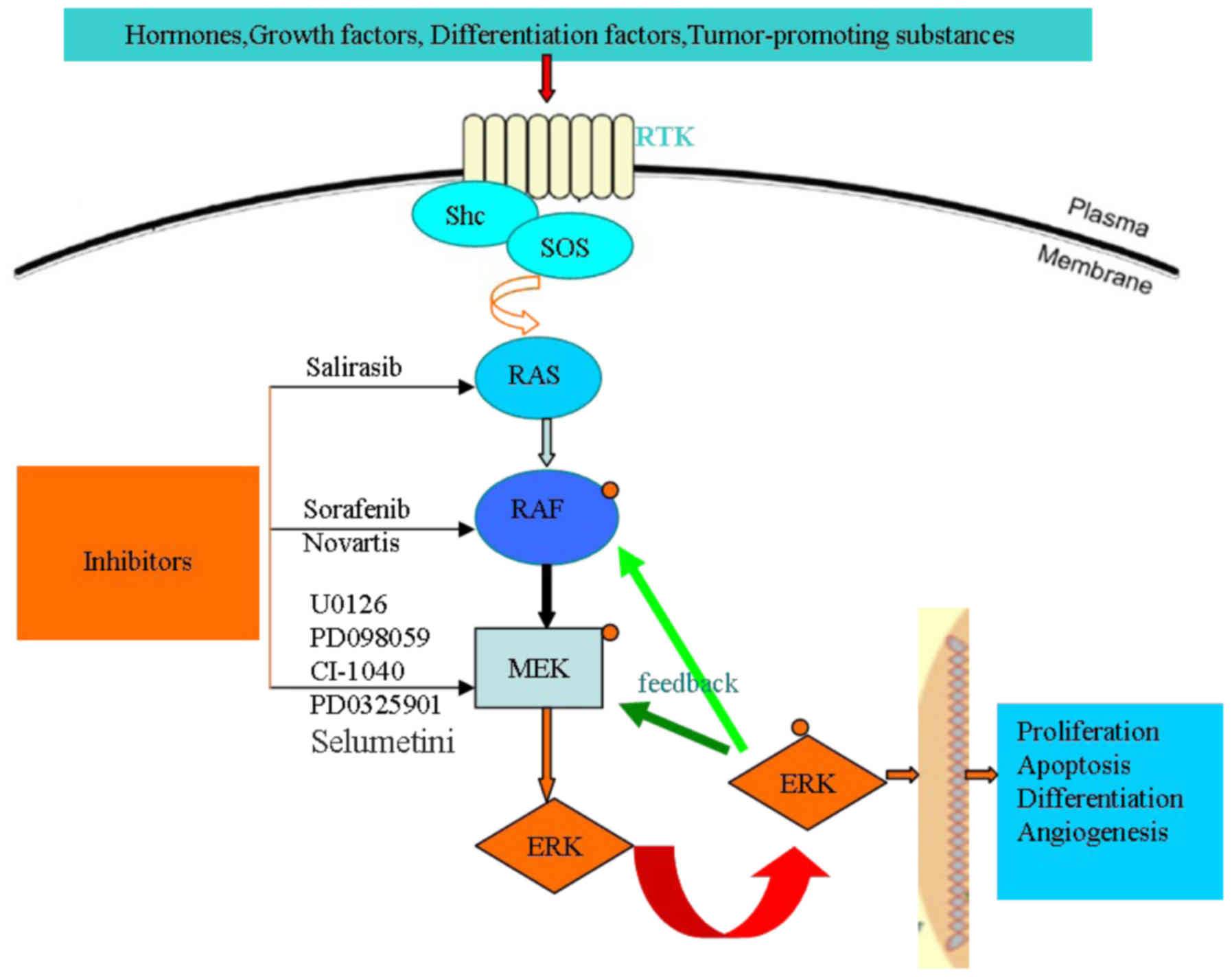
224. Kolch W. Coordinating ERK/MAPK signalling through scaffolds and inhibitors. Nat Rev Mol Cell Biol. 2005;6:827-37 [PubMed] [Google Scholar]
225. Levchenko A, Bruck J, Sternberg PW. Scaffold proteins may biphasically affect the levels of mitogen-activated protein kinase signaling and reduce its threshold properties. Proc Natl Acad Sci U S A. 2000;97:5818-23 [PMC free article] [PubMed] [Google Scholar]
226. Casar B, Arozarena I, Sanz-Moreno V, et al. Ras subcellular localization defines extracellular signal-regulated kinase 1 and 2 substrate specificity through distinct utilization of scaffold proteins. Mol Cell Biol. 2009;29:1338-53 [PMC free article] [PubMed] [Google Scholar]
227. Casar B, Pinto A, Crespo P. Essential role of ERK dimers in the activation of cytoplasmic but not nuclear substrates by ERK-scaffold complexes. Mol Cell. 2008;31:708-21 [PubMed] [Google Scholar]
228. Calvo F, Agudo-Ibanez L, Crespo P. The Ras-ERK pathway: understanding site-specific signaling provides hope of new anti-tumor therapies. Bioessays. 2010;32:412-21 [PubMed] [Google Scholar]
229. Dard N, Peter M. Scaffold proteins in MAP kinase signaling: more than simple passive activating platforms. Bioessays. 2006;28:146-56 [PubMed] [Google Scholar]
230. Brown MD, Sacks DB. Protein scaffolds in MAP kinase signalling. Cell Signal. 2009;21:462-9 [PMC free article] [PubMed] [Google Scholar]
231. Casar B, Pinto A, Crespo P. ERK dimers and scaffold proteins: unexpected partners for a forgotten (cytoplasmic) task. Cell Cycle. 2009;8:1007-13 [PubMed] [Google Scholar]
232. Wimmer R, Baccarini M. Partner exchange: protein-protein interactions in the Raf pathway. Trends Biochem Sci. 2010;35:660-8 [PubMed] [Google Scholar]
233. Morrison DK. KSR: a MAPK scaffold of the Ras pathway?J Cell Sci. 2001;114:1609-12 [PubMed] [Google Scholar]
234. Claperon A, Therrien M. KSR and CNK: two scaffolds regulating RAS-mediated RAF activation. Oncogene. 2007;26:3143-58 [PubMed] [Google Scholar]
235. Therrien M, Chang HC, Solomon NM, Karim FD, Wassarman DA, Rubin GM. KSR, a novel protein kinase required for RAS signal transduction. Cell. 1995;83:879-88 [PubMed] [Google Scholar]
236. Kornfeld K, Hom DB, Horvitz HR. The ksr-1 gene encodes a novel protein kinase involved in Ras-mediated signaling in C. elegans. Cell. 1995;83:903-13 [PubMed] [Google Scholar]
237. Sundaram M, Han M. The C. elegans ksr-1 gene encodes a novel Raf-related kinase involved in Ras-mediated signal transduction. Cell. 1995;83:889-901 [PubMed] [Google Scholar]
238. Kolesnick R, Xing HR. Inflammatory bowel disease reveals the kinase activity of KSR1. J Clin Invest. 2004;114:1233-7 [PMC free article] [PubMed] [Google Scholar]
239. Boudeau J, Miranda-Saavedra D, Barton GJ, Alessi DR. Emerging roles of pseudokinases. Trends Cell Biol. 2006;16:443-52 [PubMed] [Google Scholar]
240. Therrien M, Michaud NR, Rubin GM, Morrison DK. KSR modulates signal propagation within the MAPK cascade. Genes Dev. 1996;10:2684-95 [PubMed] [Google Scholar]
241. McKay MM, Ritt DA, Morrison DK. Signaling dynamics of the KSR1 scaffold complex. Proc Natl Acad Sci U S A. 2009;106:11022-7 [PMC free article] [PubMed] [Google Scholar]
242. Joneson T, Fulton JA, Volle DJ, Chaika OV, Bar-Sagi D, Lewis RE. Kinase suppressor of Ras inhibits the activation of extracellular ligand-regulated (ERK) mitogen-activated protein (MAP) kinase by growth factors, activated Ras, and Ras effectors. J Biol Chem. 1998;273:7743-8 [PubMed] [Google Scholar]
243. Ohmachi M, Rocheleau CE, Church D, Lambie E, Schedl T, Sundaram MV. C. elegans ksr-1 and ksr-2 have both unique and redundant functions and are required for MPK-1 ERK phosphorylation. Curr Biol. 2002;12:427-33 [PubMed] [Google Scholar]
244. Kortum RL, Costanzo DL, Haferbier J, et al. The molecular scaffold kinase suppressor of Ras 1 (KSR1) regulates adipogenesis. Mol Cell Biol. 2005;25:7592-604 [PMC free article] [PubMed] [Google Scholar]
245. Kortum RL, Lewis RE. The molecular scaffold KSR1 regulates the proliferative and oncogenic potential of cells. Mol Cell Biol. 2004;24:4407-16 [PMC free article] [PubMed] [Google Scholar]
246. Kortum RL, Johnson HJ, Costanzo DL, et al. The molecular scaffold kinase suppressor of Ras 1 is a modifier of RasV12-induced and replicative senescence. Mol Cell Biol. 2006;26:2202-14 [PMC free article] [PubMed] [Google Scholar]
247. Nguyen A, Burack WR, Stock JL, et al. Kinase suppressor of Ras (KSR) is a scaffold which facilitates mitogen-activated protein kinase activation in vivo. Mol Cell Biol. 2002;22:3035-45 [PMC free article] [PubMed] [Google Scholar]
248. Lozano J, Xing R, Cai Z, et al. Deficiency of kinase suppressor of Ras1 prevents oncogenic ras signaling in mice. Cancer Res. 2003;63:4232-8 [PubMed] [Google Scholar]
249. Xing HR, Cordon-Cardo C, Deng X, et al. Pharmacologic inactivation of kinase suppressor of ras-1 abrogates Ras-mediated pancreatic cancer. Nat Med. 2003;9:1266-8 [PubMed] [Google Scholar]
250. Kim M, Yan Y, Kortum RL, et al. Expression of kinase suppressor of Ras1 enhances cisplatin-induced extracellular signal-regulated kinase activation and cisplatin sensitivity. Cancer Res. 2005;65:3986-92 [PubMed] [Google Scholar]
251. Stoeger SM, Cowan KH. Characterization of kinase suppressor of Ras-1 expression and anticancer drug sensitivity in human cancer cell lines. Cancer Chemother Pharmacol. 2009;63:807-18 [PubMed] [Google Scholar]
252. Dougherty MK, Ritt DA, Zhou M, et al. KSR2 is a calcineurin substrate that promotes ERK cascade activation in response to calcium signals. Mol Cell. 2009;34:652-62 [PMC free article] [PubMed] [Google Scholar]
253. Liu L, Channavajhala PL, Rao VR, et al. Proteomic characterization of the dynamic KSR-2 interactome, a signaling scaffold complex in MAPK pathway. Biochim Biophys Acta. 2009;1794:1485-95 [PubMed] [Google Scholar]
254. Costanzo-Garvey DL, Pfluger PT, Dougherty MK, et al. KSR2 is an essential regulator of AMP kinase, energy expenditure, and insulin sensitivity. Cell Metab. 2009;10:366-78 [PMC free article] [PubMed] [Google Scholar]
255. Revelli JP, Smith D, Allen J, et al. Profound obesity secondary to hyperphagia in mice lacking kinase suppressor of Ras 2. Obesity (Silver Spring). Epub 2010 Dec 2 [PubMed] [Google Scholar]
256. Therrien M, Wong AM, Rubin GM. CNK, a RAF-binding multidomain protein required for RAS signaling. Cell. 1998;95:343-53 [PubMed] [Google Scholar]
257. Douziech M, Sahmi M, Laberge G, Therrien M. A KSR/CNK complex mediated by HYP, a novel SAM domain-containing protein, regulates RAS-dependent RAF activation in Drosophila. Genes Dev. 2006;20:807-19 [PMC free article] [PubMed] [Google Scholar]
258. Roignant JY, Hamel S, Janody F, Treisman JE. The novel SAM domain protein Aveugle is required for Raf activation in the Drosophila EGF receptor signaling pathway. Genes Dev. 2006;20:795-806 [PMC free article] [PubMed] [Google Scholar]
259. Bumeister R, Rosse C, Anselmo A, Camonis J, White MA. CNK2 couples NGF signal propagation to multiple regulatory cascades driving cell differentiation. Curr Biol. 2004;14:439-45 [PubMed] [Google Scholar]
260. Ziogas A, Moelling K, Radziwill G. CNK1 is a scaffold protein that regulates Src-mediated Raf-1 activation. J Biol Chem. 2005;280:24205-11 [PubMed] [Google Scholar]
261. Rabizadeh S, Xavier RJ, Ishiguro K, et al. The scaffold protein CNK1 interacts with the tumor suppressor RASSF1A and augments RASSF1A-induced cell death. J Biol Chem. 2004;279:29247-54 [PubMed] [Google Scholar]
262. Briggs MW, Sacks DB. IQGAP proteins are integral components of cytoskeletal regulation. EMBO Rep. 2003;4:571-4 [PMC free article] [PubMed] [Google Scholar]
263. White CD, Brown MD, Sacks DB. IQGAPs in cancer: a family of scaffold proteins underlying tumorigenesis. FEBS Lett. 2009;583:1817-24 [PMC free article] [PubMed] [Google Scholar]
264. Roy M, Li Z, Sacks DB. IQGAP1 binds ERK2 and modulates its activity. J Biol Chem. 2004;279:17329-37 [PubMed] [Google Scholar]
265. Roy M, Li Z, Sacks DB. IQGAP1 is a scaffold for mitogen-activated protein kinase signaling. Mol Cell Biol. 2005;25:7940-52 [PMC free article] [PubMed] [Google Scholar]
266. Ren JG, Li Z, Sacks DB. IQGAP1 modulates activation of B-Raf. Proc Natl Acad Sci U S A. 2007;104:10465-9 [PMC free article] [PubMed] [Google Scholar]
267. Takemoto H, Doki Y, Shiozaki H, et al. Localization of IQGAP1 is inversely correlated with intercellular adhesion mediated by e-cadherin in gastric cancers. Int J Cancer. 2001;91:783-8 [PubMed] [Google Scholar]
268. Jadeski L, Mataraza JM, Jeong HW, Li Z, Sacks DB. IQGAP1 stimulates proliferation and enhances tumorigenesis of human breast epithelial cells. J Biol Chem. 2008;283:1008-17 [PubMed] [Google Scholar]
269. Rajalingam K, Rudel T. Ras-Raf signaling needs prohibitin. Cell Cycle. 2005;4:1503-5 [PubMed] [Google Scholar]
270. Nijtmans LG, Artal SM, Grivell LA, Coates PJ. The mitochondrial PHB complex: roles in mitochondrial respiratory complex assembly, ageing and degenerative disease. Cell Mol Life Sci. 2002;59:143-55 [PubMed] [Google Scholar]
271. Ryu JW, Kim HJ, Lee YS, et al. The proteomics approach to find biomarkers in gastric cancer. J Korean Med Sci. 2003;18:505-9 [PMC free article] [PubMed] [Google Scholar]
272. Spurdle AB, Hopper JL, Chen X, et al. Prohibitin 3′ untranslated region polymorphism and breast cancer risk in Australian women. Lancet. 2002;360:925-6 [PubMed] [Google Scholar]
273. Torii S, Kusakabe M, Yamamoto T, Maekawa M, Nishida E. Sef is a spatial regulator for Ras/MAP kinase signaling. Dev Cell. 2004;7:33-44 [PubMed] [Google Scholar]
274. Darby S, Sahadevan K, Khan MM, Robson CN, Leung HY, Gnanapragasam VJ. Loss of Sef (similar expression to FGF) expression is associated with high grade and metastatic prostate cancer. Oncogene. 2006;25:4122-7 [PubMed] [Google Scholar]
275. DeFea KA, Zalevsky J, Thoma MS, Dery O, Mullins RD, Bunnett NW. beta-arrestin-dependent endocytosis of proteinase-activated receptor 2 is required for intracellular targeting of activated ERK1/2. J Cell Biol. 2000;148:1267-81 [PMC free article] [PubMed] [Google Scholar]
276. Shenoy SK, Lefkowitz RJ. Receptor-specific ubiquitination of beta-arrestin directs assembly and targeting of seven-transmembrane receptor signalosomes. J Biol Chem. 2005;280:15315-24 [PubMed] [Google Scholar]
277. Teis D, Wunderlich W, Huber LA. Localization of the MP1-MAPK scaffold complex to endosomes is mediated by p14 and required for signal transduction. Dev Cell. 2002;3:803-14 [PubMed] [Google Scholar]
278. Teis D, Taub N, Kurzbauer R, et al. p14-MP1-MEK1 signaling regulates endosomal traffic and cellular proliferation during tissue homeostasis. J Cell Biol. 2006;175:861-8 [PMC free article] [PubMed] [Google Scholar]
279. Nada S, Hondo A, Kasai A, et al. The novel lipid raft adaptor p18 controls endosome dynamics by anchoring the MEK-ERK pathway to late endosomes. EMBO J. 2009;28:477-89 [PMC free article] [PubMed] [Google Scholar]
280. Schaeffer HJ, Catling AD, Eblen ST, Collier LS, Krauss A, Weber MJ. MP1: a MEK binding partner that enhances enzymatic activation of the MAP kinase cascade. Science. 1998;281:1668-71 [PubMed] [Google Scholar]
281. Pullikuth A, McKinnon E, Schaeffer HJ, Catling AD. The MEK1 scaffolding protein MP1 regulates cell spreading by integrating PAK1 and Rho signals. Mol Cell Biol. 2005;25:5119-33 [PMC free article] [PubMed] [Google Scholar]
282. Okamoto I, Pirker C, Bilban M, et al. Seven novel and stable translocations associated with oncogenic gene expression in malignant melanoma. Neoplasia. 2005;7:303-11 [PMC free article] [PubMed] [Google Scholar]
283. Sharma C, Vomastek T, Tarcsafalvi A, et al. MEK partner 1 (MP1): regulation of oligomerization in MAP kinase signaling. J Cell Biochem. 2005;94:708-19 [PubMed] [Google Scholar]
284. Vomastek T, Schaeffer H-J, Tarcsafalvi A, Smolkin ME, Bissonette EA, Weber MJ. Modular construction of a signaling scaffold: MORG1 interacts with components of the ERK cascade and links ERK signaling to specific agonists. Proc Natl Acad Sci U S A. 2004;101:6981-6 [PMC free article] [PubMed] [Google Scholar]
285. Hopfer U, Hopfer H, Jablonski K, Stahl RAK, Wolf G. The novel WD-repeat protein Morg1 acts as a molecular scaffold for hypoxia-inducible factor prolyl hydroxylase 3 (PHD3). J Biol Chem. 2006;281:8645-55 [PubMed] [Google Scholar]
286. Deakin NO, Turner CE. Paxillin comes of age. J Cell Sci. 2008;121:2435-44 [PMC free article] [PubMed] [Google Scholar]
287. Ishibe S, Joly D, Zhu X, Cantley LG. Phosphorylation-dependent paxillin-ERK association mediates hepatocyte growth factor-stimulated epithelial morphogenesis. Mol Cell. 2003;12:1275-85 [PubMed] [Google Scholar]
288. Hagel M, George EL, Kim A, et al. The adaptor protein paxillin is essential for normal development in the mouse and is a critical transducer of fibronectin signaling. Mol Cell Biol. 2002;22:901-15 [PMC free article] [PubMed] [Google Scholar]
289. Brown MC, Turner CE. Paxillin: adapting to change. Physiol Rev. 2004;84:1315-39 [PubMed] [Google Scholar]
290. Azuma K, Tanaka M, Uekita T, et al. Tyrosine phosphorylation of paxillin affects the metastatic potential of human osteosarcoma. Oncogene. 2005;24:4754-64 [PubMed] [Google Scholar]
291. Li D, Ding J, Wang X, Wang C, Wu T. Fibronectin promotes tyrosine phosphorylation of paxillin and cell invasiveness in the gastric cancer cell line AGS. Tumori. 2009;95:769-79 [PubMed] [Google Scholar]
292. Trakul N, Rosner MR. Modulation of the MAP kinase signaling cascade by Raf kinase inhibitory protein. Cell Res. 2005;15:19-23 [PubMed] [Google Scholar]
293. Keller ET, Fu Z, Brennan M. The biology of a prostate cancer metastasis suppressor protein: Raf kinase inhibitor protein. J Cell Biochem. 2005;94:273-8 [PubMed] [Google Scholar]
294. Granovsky AE, Rosner MR. Raf kinase inhibitory protein: a signal transduction modulator and metastasis suppressor. Cell Res. 2008;18:452-7 [PubMed] [Google Scholar]
295. Trakul N, Menard RE, Schade GR, Qian Z, Rosner MR. Raf kinase inhibitory protein regulates Raf-1 but not B-Raf kinase activation. J Biol Chem. 2005;280:24931-40 [PubMed] [Google Scholar]
296. Park S, Yeung ML, Beach S, Shields JM, Yeung KC. RKIP downregulates B-Raf kinase activity in melanoma cancer cells. Oncogene. 2005;24:3535-40 [PubMed] [Google Scholar]
297. Shin SY, Rath O, Choo SM, et al. Positive- and negative-feedback regulations coordinate the dynamic behavior of the Ras-Raf-MEK-ERK signal transduction pathway. J Cell Sci. 2009;122:425-35 [PubMed] [Google Scholar]
298. Fu Z, Smith PC, Zhang L, et al. Effects of raf kinase inhibitor protein expression on suppression of prostate cancer metastasis. J Natl Cancer Inst. 2003;95:878-89 [PubMed] [Google Scholar]
299. Hagan S, Al-Mulla F, Mallon E, et al. Reduction of Raf-1 kinase inhibitor protein expression correlates with breast cancer metastasis. Clin Cancer Res. 2005;11:7392-7 [PubMed] [Google Scholar]
300. Al-Mulla F, Hagan S, Behbehani AI, et al. Raf kinase inhibitor protein expression in a survival analysis of colorectal cancer patients. J Clin Oncol. 2006;24:5672-9 [PubMed] [Google Scholar]
301. Lee HC, Tian B, Sedivy JM, Wands JR, Kim M. Loss of Raf kinase inhibitor protein promotes cell proliferation and migration of human hepatoma cells. Gastroenterology. 2006;131:1208-17 [PMC free article] [PubMed] [Google Scholar]
302. Torii S, Nakayama K, Yamamoto T, Nishida E. Regulatory mechanisms and function of ERK MAP kinases. J Biochem. 2004;136:557-61 [PubMed] [Google Scholar]
303. Lo TL, Fong CW, Yusoff P, et al. Sprouty and cancer: the first terms report. Cancer Lett. 2006;242:141-50 [PubMed] [Google Scholar]
304. Mason JM, Morrison DJ, Basson MA, Licht JD. Sprouty proteins: multifaceted negative-feedback regulators of receptor tyrosine kinase signaling. Trends Cell Biol. 2006;16:45-54 [PubMed] [Google Scholar]
305. Cabrita MA, Christofori G. Sprouty proteins, masterminds of receptor tyrosine kinase signaling. Angiogenesis. 2008;11:53-62 [PubMed] [Google Scholar]
306. Guy GR, Jackson RA, Yusoff P, Chow SY. Sprouty proteins: modified modulators, matchmakers or missing links?J Endocrinol. 2009;203:191-202 [PubMed] [Google Scholar]
307. Hanafusa H, Torii S, Yasunaga T, Matsumoto K, Nishida E. Shp2, an SH2-containing protein-tyrosine phosphatase, positively regulates receptor tyrosine kinase signaling by dephosphorylating and inactivating the inhibitor Sprouty. J Biol Chem. 2004;279:22992-5 [PubMed] [Google Scholar]
308. Wakioka T, Sasaki A, Kato R, et al. Spred is a Sprouty-related suppressor of Ras signalling. Nature. 2001;412:647-51 [PubMed] [Google Scholar]
309. Sasaki A, Taketomi T, Kato R, et al. Mammalian Sprouty4 suppresses Ras-independent ERK activation by binding to Raf1. Nat Cell Biol. 2003;5:427-32 [PubMed] [Google Scholar]
310. Kwabi-Addo B, Wang J, Erdem H, et al. The expression of Sprouty1, an inhibitor of fibroblast growth factor signal transduction, is decreased in human prostate cancer. Cancer Res. 2004;64:4728-35 [PubMed] [Google Scholar]
311. Fong CW, Chua MS, McKie AB, et al. Sprouty 2, an inhibitor of mitogen-activated protein kinase signaling, is down-regulated in hepatocellular carcinoma. Cancer Res. 2006;66:2048-58 [PubMed] [Google Scholar]
312. Niculescu-Duvaz D, Niculescu-Duvaz I, Suijkerbuijk BM, et al. Novel tricyclic pyrazole BRAF inhibitors with imidazole or furan central scaffolds. Bioorg Med Chem. 2010;18:6934-52 [PMC free article] [PubMed] [Google Scholar]
313. Zebisch A, Troppmair J. Back to the roots: the remarkable RAF oncogene story. Cell Mol Life Sci. 2006;63:1314-30 [PubMed] [Google Scholar]
314. Zebisch A, Czernilovsky AP, Keri G, Smigelskaite J, Sill H, Troppmair J. Signaling through RAS-RAF-MEK-ERK: from basics to bedside. Curr Med Chem. In press [PubMed] [Google Scholar]
315. Rajagopalan H, Bardelli A, Lengauer C, Kinzler KW, Vogelstein B, Velculescu VE. Tumorigenesis: RAF/RAS oncogenes and mismatch-repair status. Nature. 2002;418:934. [PubMed] [Google Scholar]
316. Dhomen N, Reis-Filho JS, da Rocha Dias S, et al. Oncogenic Braf induces melanocyte senescence and melanoma in mice. Cancer Cell. 2009;15:294-303 [PubMed] [Google Scholar]
317. Dankort D, Curley DP, Cartlidge RA, et al. Braf(V600E) cooperates with Pten loss to induce metastatic melanoma. Nat Genet. 2009;41:544-52 [PMC free article] [PubMed] [Google Scholar]
318. Pollock PM, Harper UL, Hansen KS, et al. High frequency of BRAF mutations in nevi. Nat Genet. 2003;33:19-20 [PubMed] [Google Scholar]
319. Michaloglou C, Vredeveld LC, Soengas MS, et al. BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature. 2005;436:720-4 [PubMed] [Google Scholar]
320. Storm SM, Rapp UR. Oncogene activation: c-raf-1 gene mutations in experimental and naturally occurring tumors. Toxicol Lett. 1993;67:201-10 [PubMed] [Google Scholar]
321. Emuss V, Garnett M, Mason C, Marais R. Mutations of C-RAF are rare in human cancer because C-RAF has a low basal kinase activity compared with B-RAF. Cancer Res. 2005;65:9719-26 [PubMed] [Google Scholar]
322. Zebisch A, Staber PB, Delavar A, et al. Two transforming C-RAF germ-line mutations identified in patients with therapy-related acute myeloid leukemia. Cancer Res. 2006;66:3401-8 [PubMed] [Google Scholar]
323. Zebisch A, Haller M, Hiden K, et al. Loss of RAF kinase inhibitor protein is a somatic event in the pathogenesis of therapy-related acute myeloid leukemias with C-RAF germline mutations. Leukemia. 2009;23:1049-53 [PubMed] [Google Scholar]
324. Yu J, Deshmukh H, Gutmann RJ, et al. Alterations of BRAF and HIPK2 loci predominate in sporadic pilocytic astrocytoma. Neurology. 2009;73:1526-31 [PMC free article] [PubMed] [Google Scholar]
325. Jones DT, Kocialkowski S, Liu L, et al. Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res. 2008;68:8673-7 [PMC free article] [PubMed] [Google Scholar]
326. Jones DT, Kocialkowski S, Liu L, Pearson DM, Ichimura K, Collins VP. Oncogenic RAF1 rearrangement and a novel BRAF mutation as alternatives to KIAA1549:BRAF fusion in activating the MAPK pathway in pilocytic astrocytoma. Oncogene. 2009;28:2119-23 [PMC free article] [PubMed] [Google Scholar]
327. Ciampi R, Knauf JA, Kerler R, et al. Oncogenic AKAP9-BRAF fusion is a novel mechanism of MAPK pathway activation in thyroid cancer. J Clin Invest. 2005;115:94-101 [PMC free article] [PubMed] [Google Scholar]
328. Palanisamy N, Ateeq B, Kalyana-Sundaram S, et al. Rearrangements of the RAF kinase pathway in prostate cancer, gastric cancer and melanoma. Nat Med. 2010;16:793-8 [PMC free article] [PubMed] [Google Scholar]
329. Maldonado JL, Fridlyand J, Patel H, et al. Determinants of BRAF mutations in primary melanomas. J Natl Cancer Inst. 2003;95:1878-90 [PubMed] [Google Scholar]
330. Kawakami T, Okamoto K, Sugihara H, et al. The roles of supernumerical X chromosomes and XIST expression in testicular germ cell tumors. J Urol. 2003;169:1546-52 [PubMed] [Google Scholar]
331. Aoki Y, Niihori T, Narumi Y, Kure S, Matsubara Y. The RAS/MAPK syndromes: novel roles of the RAS pathway in human genetic disorders. Hum Mutat. 2008;29:992-1006 [PubMed] [Google Scholar]
332. Denayer E, de Ravel T, Legius E. Clinical and molecular aspects of RAS related disorders. J Med Genet. 2008;45:695-703 [PubMed] [Google Scholar]
333. Schubbert S, Shannon K, Bollag G. Hyperactive Ras in developmental disorders and cancer. Nat Rev Cancer. 2007;7:295-308 [PubMed] [Google Scholar]
334. Niihori T, Aoki Y, Narumi Y, et al. Germline KRAS and BRAF mutations in cardio-facio-cutaneous syndrome. Nat Genet. 2006;38:294-6 [PubMed] [Google Scholar]
335. Rodriguez-Viciana P, Tetsu O, Tidyman WE, et al. Germline mutations in genes within the MAPK pathway cause cardio-facio-cutaneous syndrome. Science. 2006;311:1287-90 [PubMed] [Google Scholar]
336. Rodriguez-Viciana P, Rauen KA. Biochemical characterization of novel germline BRAF and MEK mutations in cardio-facio-cutaneous syndrome. Methods Enzymol. 2008;438:277-89 [PubMed] [Google Scholar]
337. Sarkozy A, Carta C, Moretti S, et al. Germline BRAF mutations in Noonan, LEOPARD, and cardiofaciocutaneous syndromes: molecular diversity and associated phenotypic spectrum. Hum Mutat. 2009;30:695-702 [PMC free article] [PubMed] [Google Scholar]
338. Pandit B, Sarkozy A, Pennacchio LA, et al. Gain-of-function RAF1 mutations cause Noonan and LEOPARD syndromes with hypertrophic cardiomyopathy. Nat Genet. 2007;39:1007-12 [PubMed] [Google Scholar]
339. Razzaque MA, Nishizawa T, Komoike Y, et al. Germline gain-of-function mutations in RAF1 cause Noonan syndrome. Nat Genet. 2007;39:1013-7 [PubMed] [Google Scholar]
340. Kobayashi T, Aoki Y, Niihori T, et al. Molecular and clinical analysis of RAF1 in Noonan syndrome and related disorders: dephosphorylation of serine 259 as the essential mechanism for mutant activation. Hum Mutat. 2010;31:284-94 [PubMed] [Google Scholar]
341. Laux D, Kratz C, Sauerbrey A. Common acute lymphoblastic leukemia in a girl with genetically confirmed LEOPARD syndrome. J Pediatr Hematol Oncol. 2008;30:602-4 [PubMed] [Google Scholar]
342. Merks JH, Caron HN, Hennekam RC. High incidence of malformation syndromes in a series of 1,073 children with cancer. Am J Med Genet A. 2005;134A:132-43 [PubMed] [Google Scholar]
343. Ohtake A, Aoki Y, Saito Y, et al. Non-Hodgkin lymphoma in a patient with cardiofaciocutaneous syndrome. J Pediatr Hematol Oncol. Epub 2010 Jun 2 [PubMed] [Google Scholar]
344. Ucar C, Calyskan U, Martini S, Heinritz W. Acute myelomonocytic leukemia in a boy with LEOPARD syndrome (PTPN11 gene mutation positive). J Pediatr Hematol Oncol. 2006;28:123-5 [PubMed] [Google Scholar]
345. van Den Berg H, Hennekam RC. Acute lymphoblastic leukaemia in a patient with cardiofaciocutaneous syndrome. J Med Genet. 1999;36:799-800 [PMC free article] [PubMed] [Google Scholar]
346. Inamdar GS, Madhunapantula SV, Robertson GP. Targeting the MAPK pathway in melanoma: why some approaches succeed and other fail. Biochem Pharmacol. 2010;80:624-37 [PMC free article] [PubMed] [Google Scholar]
347. Mendel DB, Laird AD, Xin X, et al. In vivo antitumor activity of SU11248, a novel tyrosine kinase inhibitor targeting vascular endothelial growth factor and platelet-derived growth factor receptors: determination of a pharmacokinetic/pharmacodynamic relationship. Clin Cancer Res. 2003;9:327-37 [PubMed] [Google Scholar]
348. Montagut C, Settleman J. Targeting the RAF-MEK-ERK pathway in cancer therapy. Cancer Lett. 2009;283:125-34 [PubMed] [Google Scholar]
349. Dhomen N, Marais R. BRAF signaling and targeted therapies in melanoma. Hematol Oncol Clin North Am. 2009;23:529-45, ix [PubMed] [Google Scholar]
350. Eisen T, Ahmad T, Flaherty KT, et al. Sorafenib in advanced melanoma: a phase II randomised discontinuation trial analysis. Br J Cancer. 2006;95:581-6 [PMC free article] [PubMed] [Google Scholar]
351. Wilhelm SM, Carter C, Tang L, et al. BAY 43-9006 exhibits broad spectrum oral antitumor activity and targets the RAF/MEK/ERK pathway and receptor tyrosine kinases involved in tumor progression and angiogenesis. Cancer Res. 2004;64:7099-109 [PubMed] [Google Scholar]
352. Roberts PJ, Der CJ. Targeting the Raf-MEK-ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene. 2007;26:3291-310 [PubMed] [Google Scholar]
Ras And Raf
353. Sondergaard JN, Nazarian R, Wang Q, et al. Differential sensitivity of melanoma cell lines with BRAFV600E mutation to the specific Raf inhibitor PLX4032. J Transl Med. 2010;8:39. [PMC free article] [PubMed] [Google Scholar]
354. Tsai J, Lee JT, Wang W, et al. Discovery of a selective inhibitor of oncogenic B-Raf kinase with potent antimelanoma activity. Proc Natl Acad Sci U S A. 2008;105:3041-6 [PMC free article] [PubMed] [Google Scholar]
355. Flaherty KT, Puzanov I, Kim KB, et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med. 2010;363:809-19 [PMC free article] [PubMed] [Google Scholar]
356. Khazak V, Astsaturov I, Serebriiskii IG, Golemis EA. Selective Raf inhibition in cancer therapy. Expert Opin Ther Targets. 2007;11:1587-609 [PMC free article] [PubMed] [Google Scholar]
357. Davis RK, Chellappan S. Disrupting the Rb-Raf-1 interaction: a potential therapeutic target for cancer. Drug News Perspect. 2008;21:331-5 [PMC free article] [PubMed] [Google Scholar]
358. Cunningham CC, Holmlund JT, Schiller JH, et al. A phase I trial of c-Raf kinase antisense oligonucleotide ISIS 5132 administered as a continuous intravenous infusion in patients with advanced cancer. Clin Cancer Res. 2000;6:1626-31 [PubMed] [Google Scholar]
359. Rudin CM, Marshall JL, Huang CH, et al. Delivery of a liposomal c-raf-1 antisense oligonucleotide by weekly bolus dosing in patients with advanced solid tumors: a phase I study. Clin Cancer Res. 2004;10:7244-51 [PubMed] [Google Scholar]
360. Dritschilo A, Huang CH, Rudin CM, et al. Phase I study of liposome-encapsulated c-raf antisense oligodeoxyribonucleotide infusion in combination with radiation therapy in patients with advanced malignancies. Clin Cancer Res. 2006;12:1251-9 [PubMed] [Google Scholar]
361. Shim MS, Kwon YJ. Efficient and targeted delivery of siRNA in vivo. FEBS J. 2010;277:4814-27 [PubMed] [Google Scholar]
362. Sharma A, Trivedi NR, Zimmerman MA, Tuveson DA, Smith CD, Robertson GP. Mutant V599EB-Raf regulates growth and vascular development of malignant melanoma tumors. Cancer Res. 2005;65:2412-21 [PubMed] [Google Scholar]
363. Tran MA, Gowda R, Sharma A, et al. Targeting V600EB-Raf and Akt3 using nanoliposomal-small interfering RNA inhibits cutaneous melanocytic lesion development. Cancer Res. 2008;68:7638-49 [PMC free article] [PubMed] [Google Scholar]
364. Leng Q, Scaria P, Lu P, Woodle MC, Mixson AJ. Systemic delivery of HK Raf-1 siRNA polyplexes inhibits MDA-MB-435 xenografts. Cancer Gene Ther. 2008;15:485-95 [PMC free article] [PubMed] [Google Scholar]
365. Karreth FA, DeNicola GM, Winter SP, Tuveson DA. C-Raf inhibits MAPK activation and transformation by B-Raf(V600E). Mol Cell. 2009;36:477-86 [PubMed] [Google Scholar]
366. Daouti S, Higgins B, Kolinsky K, et al. Preclinical in vivo evaluation of efficacy, pharmacokinetics, and pharmacodynamics of a novel MEK1/2 kinase inhibitor RO5068760 in multiple tumor models. Mol Cancer Ther. 2010;9:134-44 [PubMed] [Google Scholar]
367. Solit DB, Garraway LA, Pratilas CA, et al. BRAF mutation predicts sensitivity to MEK inhibition. Nature. 2006;439:358-62 [PMC free article] [PubMed] [Google Scholar]
368. Lorusso PM, Adjei AA, Varterasian M, et al. Phase I and pharmacodynamic study of the oral MEK inhibitor CI-1040 in patients with advanced malignancies. J Clin Oncol. 2005;23:5281-93 [PubMed] [Google Scholar]
369. Rinehart J, Adjei AA, Lorusso PM, et al. Multicenter phase II study of the oral MEK inhibitor, CI-1040, in patients with advanced non-small-cell lung, breast, colon, and pancreatic cancer. J Clin Oncol. 2004;22:4456-62 [PubMed] [Google Scholar]
370. Fremin C, Meloche S. From basic research to clinical development of MEK1/2 inhibitors for cancer therapy. J Hematol Oncol. 2010;3:8. [PMC free article] [PubMed] [Google Scholar]
371. Santos SD, Verveer PJ, Bastiaens PI. Growth factor-induced MAPK network topology shapes Erk response determining PC-12 cell fate. Nat Cell Biol. 2007;9:324-30 [PubMed] [Google Scholar]
372. Schilling M, Maiwald T, Hengl S, et al. Theoretical and experimental analysis links isoform-specific ERK signalling to cell fate decisions. Mol Syst Biol. 2009;5:334. [PMC free article] [PubMed] [Google Scholar]
373. Orton RJ, Sturm OE, Vyshemirsky V, Calder M, Gilbert DR, Kolch W. Computational modelling of the receptor-tyrosine-kinase-activated MAPK pathway. Biochem J. 2005;392:249-61 [PMC free article] [PubMed] [Google Scholar]
374. Fujioka A, Terai K, Itoh RE, et al. Dynamics of the Ras/ERK MAPK cascade as monitored by fluorescent probes. J Biol Chem. 2006;281:8917-26 [PubMed] [Google Scholar]
375. Chen WW, Schoeberl B, Jasper PJ, et al. Input-output behavior of ErbB signaling pathways as revealed by a mass action model trained against dynamic data. Mol Syst Biol. 2009;5:239. [PMC free article] [PubMed] [Google Scholar]
376. Scheele JS, Rhee JM, Boss GR. Determination of absolute amounts of GDP and GTP bound to Ras in mammalian cells: comparison of parental and Ras-overproducing NIH 3T3 fibroblasts. Proc Natl Acad Sci U S A. 1995;92:1097-100 [PMC free article] [PubMed] [Google Scholar]
377. Ferrell JE., Jr How responses get more switch-like as you move down a protein kinase cascade. Trends Biochem Sci. 1997;22:288-9 [PubMed] [Google Scholar]
378. Brown GC, Hoek JB, Kholodenko BN. Why do protein kinase cascades have more than one level?Trends Biochem Sci. 1997;22:288. [PubMed] [Google Scholar]
379. Huang CY, Ferrell JE., Jr Ultrasensitivity in the mitogen-activated protein kinase cascade. Proc Natl Acad Sci U S A. 1996;93:10078-83 [PMC free article] [PubMed] [Google Scholar]
380. Qiao L, Nachbar RB, Kevrekidis IG, Shvartsman SY. Bistability and oscillations in the Huang-Ferrell model of MAPK signaling. PLoS Comput Biol. 2007;3:1819-26 [PMC free article] [PubMed] [Google Scholar]
381. Markevich NI, Hoek JB, Kholodenko BN. Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades. J Cell Biol. 2004;164:353-9 [PMC free article] [PubMed] [Google Scholar]
382. Ferrell JE, Jr., Bhatt RR. Mechanistic studies of the dual phosphorylation of mitogen-activated protein kinase. J Biol Chem. 1997;272:19008-16 [PubMed] [Google Scholar]
383. Zhao Y, Zhang ZY. The mechanism of dephosphorylation of extracellular signal-regulated kinase 2 by mitogen-activated protein kinase phosphatase 3. J Biol Chem. 2001;276:32382-91 [PubMed] [Google Scholar]
384. Kholodenko BN, Birtwistle MR. Four-dimensional dynamics of MAPK information processing systems. Wiley Interdiscip Rev Syst Biol Med. 2009;1:28-44 [PMC free article] [PubMed] [Google Scholar]
385. Sturm OE, Orton R, Grindlay J, et al. The mammalian MAPK/ERK pathway exhibits properties of a negative feedback amplifier. Sci Signal. 2010;3:ra90. [PubMed] [Google Scholar]
386. Nakayama K, Satoh T, Igari A, Kageyama R, Nishida E. FGF induces oscillations of Hes1 expression and Ras/ERK activation. Curr Biol. 2008;18:R332-4 [PubMed] [Google Scholar]
387. Shankaran H, Ippolito DL, Chrisler WB, et al. Rapid and sustained nuclear-cytoplasmic ERK oscillations induced by epidermal growth factor. Mol Syst Biol. 2009;5:332. [PMC free article] [PubMed] [Google Scholar]
388. Weber TJ, Shankaran H, Wiley HS, Opresko LK, Chrisler WB, Quesenberry RD. Basic fibroblast growth factor regulates persistent ERK oscillations in premalignant but not malignant JB6 cells. J Invest Dermatol. 2010;130:1444-56 [PubMed] [Google Scholar]
389. Kamioka Y, Yasuda S, Fujita Y, Aoki K, Matsuda M. Multiple decisive phosphorylation sites for the negative feedback regulation of SOS1 via ERK. J Biol Chem. 2010;285:33540-8 [PMC free article] [PubMed] [Google Scholar]
390. Northwood IC, Gonzalez FA, Wartmann M, Raden DL, Davis RJ. Isolation and characterization of two growth factor-stimulated protein kinases that phosphorylate the epidermal growth factor receptor at threonine 669. J Biol Chem. 1991;266:15266-76 [PubMed] [Google Scholar]
391. Sanghera JS, Hall FL, Warburton D, Campbell D, Pelech SL. Identification of epidermal growth factor Thr-669 phosphorylation site peptide kinases as distinct MAP kinases and p34cdc2. Biochim Biophys Acta. 1992;1135:335-42 [PubMed] [Google Scholar]
392. Gan Y, Shi C, Inge L, Hibner M, Balducci J, Huang Y. Differential roles of ERK and Akt pathways in regulation of EGFR-mediated signaling and motility in prostate cancer cells. Oncogene. 2010;29:4947-58 [PubMed] [Google Scholar]
393. Lehr S, Kotzka J, Avci H, et al. Identification of major ERK-related phosphorylation sites in Gab1. Biochemistry. 2004;43:12133-40 [PubMed] [Google Scholar]
394. Brummer T, Naegele H, Reth M, Misawa Y. Identification of novel ERK-mediated feedback phosphorylation sites at the C-terminus of B-Raf. Oncogene. 2003;22:8823-34 [PubMed] [Google Scholar]
395. Sauro HM, Kholodenko BN. Quantitative analysis of signaling networks. Prog Biophys Mol Biol. 2004;86:5-43 [PubMed] [Google Scholar]
396. Brondello JM, Brunet A, Pouyssegur J, McKenzie FR. The dual specificity mitogen-activated protein kinase phosphatase-1 and -2 are induced by the p42/p44MAPK cascade. J Biol Chem. 1997;272:1368-76 [PubMed] [Google Scholar]
397. Nakakuki T, Birtwistle MR, Saeki Y, et al. Ligand-specific c-Fos expression emerges from the spatiotemporal control of ErbB network dynamics. Cell. 2010;141:884-96 [PMC free article] [PubMed] [Google Scholar]
398. Nichols A, Camps M, Gillieron C, et al. Substrate recognition domains within extracellular signal-regulated kinase mediate binding and catalytic activation of mitogen-activated protein kinase phosphatase-3. J Biol Chem. 2000;275:24613-21 [PubMed] [Google Scholar]
399. Brondello JM, Pouyssegur J, McKenzie FR. Reduced MAP kinase phosphatase-1 degradation after p42/p44MAPK-dependent phosphorylation. Science. 1999;286:2514-7 [PubMed] [Google Scholar]
400. Zhang X, Pickin KA, Bose R, Jura N, Cole PA, Kuriyan J. Inhibition of the EGF receptor by binding of MIG6 to an activating kinase domain interface. Nature. 2007;450:741-4 [PMC free article] [PubMed] [Google Scholar]
401. Anastasi S, Fiorentino L, Fiorini M, et al. Feedback inhibition by RALT controls signal output by the ErbB network. Oncogene. 2003;22:4221-34 [PubMed] [Google Scholar]
402. Frosi Y, Anastasi S, Ballaro C, et al. A two-tiered mechanism of EGFR inhibition by RALT/MIG6 via kinase suppression and receptor degradation. J Cell Biol. 2010;189:557-71 [PMC free article] [PubMed] [Google Scholar]
403. Hao N, Behar M, Parnell SC, et al. A systems-biology analysis of feedback inhibition in the Sho1 osmotic-stress-response pathway. Curr Biol. 2007;17:659-67 [PubMed] [Google Scholar]
404. Behar M, Hao N, Dohlman HG, Elston TC. Mathematical and computational analysis of adaptation via feedback inhibition in signal transduction pathways. Biophys J. 2007;93:806-21 [PMC free article] [PubMed] [Google Scholar]
405. Goentoro L, Shoval O, Kirschner MW, Alon U. The incoherent feedforward loop can provide fold-change detection in gene regulation. Mol Cell. 2009;36:894-9 [PMC free article] [PubMed] [Google Scholar]
406. Ma W, Trusina A, El-Samad H, Lim WA, Tang C. Defining network topologies that can achieve biochemical adaptation. Cell. 2009;138:760-73 [PMC free article] [PubMed] [Google Scholar]
407. Formstecher E, Ramos JW, Fauquet M, et al. PEA-15 mediates cytoplasmic sequestration of ERK MAP kinase. Dev Cell. 2001;1:239-50 [PubMed] [Google Scholar]
408. von Kriegsheim A, Baiocchi D, Birtwistle M, et al. Cell fate decisions are specified by the dynamic ERK interactome. Nat Cell Biol. 2009;11:1458-64 [PMC free article] [PubMed] [Google Scholar]
409. Margarit SM, Sondermann H, Hall BE, et al. Structural evidence for feedback activation by Ras.GTP of the Ras-specific nucleotide exchange factor SOS. Cell. 2003;112:685-95 [PubMed] [Google Scholar]
410. Das J, Ho M, Zikherman J, et al. Digital signaling and hysteresis characterize ras activation in lymphoid cells. Cell. 2009;136:337-51 [PMC free article] [PubMed] [Google Scholar]
411. Smolen P, Baxter DA, Byrne JH. Bistable MAP kinase activity: a plausible mechanism contributing to maintenance of late long-term potentiation. Am J Physiol Cell Physiol. 2008;294:C503-15 [PubMed] [Google Scholar]
412. Chakraborty AK, Das J, Zikherman J, et al. Molecular origin and functional consequences of digital signaling and hysteresis during Ras activation in lymphocytes. Sci Signal. 2009;2:pt2. [PMC free article] [PubMed] [Google Scholar]
413. Joslin EJ, Shankaran H, Opresko LK, Bollinger N, Lauffenburger DA, Wiley HS. Structure of the EGF receptor transactivation circuit integrates multiple signals with cell context. Mol Biosyst. 2010;6:1293-306 [PMC free article] [PubMed] [Google Scholar]
414. Favoni RE, de Cupis A. The role of polypeptide growth factors in human carcinomas: new targets for a novel pharmacological approach. Pharmacol Rev. 2000;52:179-206 [PubMed] [Google Scholar]
415. Ruscetti FW, Akel S, Bartelmez SH. Autocrine transforming growth factor-beta regulation of hematopoiesis: many outcomes that depend on the context. Oncogene. 2005;24:5751-63 [PubMed] [Google Scholar]
416. Singh AB, Harris RC. Autocrine, paracrine and juxtacrine signaling by EGFR ligands. Cell Signal. 2005;17:1183-93 [PubMed] [Google Scholar]
417. Rodriguez-Viciana P, Warne PH, Dhand R, et al. Phosphatidylinositol-3-OH kinase as a direct target of Ras. Nature. 1994;370:527-32 [PubMed] [Google Scholar]
418. Lemmon MA. Membrane recognition by phospholipid-binding domains. Nat Rev Mol Cell Biol. 2008;9:99-111 [PubMed] [Google Scholar]
419. Kiyatkin A, Aksamitiene E, Markevich NI, Borisov NM, Hoek JB, Kholodenko BN. Scaffolding protein Grb2-associated binder 1 sustains epidermal growth factor-induced mitogenic and survival signaling by multiple positive feedback loops. J Biol Chem. 2006;281:19925-38 [PMC free article] [PubMed] [Google Scholar]
420. Montagner A, Yart A, Dance M, Perret B, Salles JP, Raynal P. A novel role for Gab1 and SHP2 in epidermal growth factor-induced Ras activation. J Biol Chem. 2005;280:5350-60 [PubMed] [Google Scholar]
421. Danial NN. BAD: undertaker by night, candyman by day. Oncogene. 2008;27Suppl 1:S53-70 [PubMed] [Google Scholar]
422. Moreto J, Llado A, Vidal-Quadras M, et al. Calmodulin modulates H-Ras mediated Raf-1 activation. Cell Signal. 2008;20:1092-103 [PubMed] [Google Scholar]
423. Alvarez-Moya B, Lopez-Alcala C, Drosten M, Bachs O, Agell N. K-Ras4B phosphorylation at Ser181 is inhibited by calmodulin and modulates K-Ras activity and function. Oncogene. 2010;29:5911-22 [PubMed] [Google Scholar]
424. Corbit KC, Trakul N, Eves EM, Diaz B, Marshall M, Rosner MR. Activation of Raf-1 signaling by protein kinase C through a mechanism involving Raf kinase inhibitory protein. J Biol Chem. 2003;278:13061-8 [PubMed] [Google Scholar]
425. Shin SY, Rath O, Zebisch A, Choo SM, Kolch W, Cho KH. Functional roles of multiple feedback loops in extracellular signal-regulated kinase and Wnt signaling pathways that regulate epithelial-mesenchymal transition. Cancer Res. 2010;70:6715-24 [PMC free article] [PubMed] [Google Scholar]
426. Houslay MD, Kolch W. Cell-type specific integration of cross-talk between extracellular signal-regulated kinase and cAMP signaling. Mol Pharmacol. 2000;58:659-68 [PubMed] [Google Scholar]
427. Yoon S, Seger R. The extracellular signal-regulated kinase: multiple substrates regulate diverse cellular functions. Growth Factors. 2006;24:21-44 [PubMed] [Google Scholar]
428. Morrison DK, Heidecker G, Rapp UR, Copeland TD. Identification of the major phosphorylation sites of the Raf-1 kinase. J Biol Chem. 1993;268:17309-16 [PubMed] [Google Scholar]
429. Stephens RM, Sithanandam G, Copeland TD, Kaplan DR, Rapp UR, Morrison DK. 95-kilodalton B-Raf serine/threonine kinase: identification of the protein and its major autophosphorylation site. Mol Cell Biol. 1992;12:3733-42 [PMC free article] [PubMed] [Google Scholar]
Articles from Genes & Cancer are provided here courtesy of Impact Journals, LLC
